You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001222_00954
You are here: Home > Sequence: MGYG000001222_00954
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | UMGS1326 sp900550775 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Monoglobales; Monoglobaceae; UMGS1326; UMGS1326 sp900550775 | |||||||||||
| CAZyme ID | MGYG000001222_00954 | |||||||||||
| CAZy Family | GH2 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 2505; End: 4652 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH2 | 23 | 651 | 3.4e-82 | 0.6316489361702128 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG3250 | LacZ | 7.76e-50 | 22 | 661 | 44 | 683 | Beta-galactosidase/beta-glucuronidase [Carbohydrate transport and metabolism]. |
| PRK10150 | PRK10150 | 5.65e-11 | 38 | 403 | 54 | 417 | beta-D-glucuronidase; Provisional |
| PRK10340 | ebgA | 2.54e-10 | 25 | 431 | 76 | 479 | cryptic beta-D-galactosidase subunit alpha; Reviewed |
| pfam00703 | Glyco_hydro_2 | 4.68e-06 | 175 | 261 | 2 | 106 | Glycosyl hydrolases family 2. This family contains beta-galactosidase, beta-mannosidase and beta-glucuronidase activities. |
| PRK09525 | lacZ | 4.97e-05 | 160 | 415 | 207 | 482 | beta-galactosidase. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| BCI62071.1 | 1.44e-224 | 7 | 707 | 27 | 750 |
| AUX47279.1 | 5.67e-223 | 6 | 678 | 22 | 704 |
| QUI21034.1 | 1.42e-219 | 15 | 713 | 18 | 738 |
| BBL06539.1 | 3.50e-215 | 6 | 675 | 11 | 699 |
| ACB77359.1 | 7.20e-201 | 22 | 699 | 35 | 743 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5N6U_A | 4.70e-54 | 7 | 601 | 33 | 654 | Crystalstructure of Beta-D-Mannosidase from Dictyoglomus thermophilum. [Dictyoglomus thermophilum H-6-12],5N6U_B Crystal structure of Beta-D-Mannosidase from Dictyoglomus thermophilum. [Dictyoglomus thermophilum H-6-12],5N6U_C Crystal structure of Beta-D-Mannosidase from Dictyoglomus thermophilum. [Dictyoglomus thermophilum H-6-12],5N6U_D Crystal structure of Beta-D-Mannosidase from Dictyoglomus thermophilum. [Dictyoglomus thermophilum H-6-12] |
| 6BYE_A | 1.84e-51 | 16 | 607 | 28 | 677 | Crystalstructure of the GH2 exo-beta-mannanase from Xanthomonas axonopodis pv. citri in complex with mannose [Xanthomonas citri pv. citri str. 306],6BYE_B Crystal structure of the GH2 exo-beta-mannanase from Xanthomonas axonopodis pv. citri in complex with mannose [Xanthomonas citri pv. citri str. 306] |
| 6BYC_A | 1.85e-51 | 16 | 607 | 28 | 677 | Crystalstructure of the GH2 exo-beta-mannanase from Xanthomonas axonopodis pv. citri [Xanthomonas citri pv. citri str. 306] |
| 6BYI_A | 1.11e-50 | 16 | 607 | 26 | 675 | Crystalstructure of the acid-base mutant (E477A) of the GH2 exo-beta-mannanase from Xanthomonas axonopodis pv. citri [Xanthomonas citri pv. citri str. 306],6BYI_B Crystal structure of the acid-base mutant (E477A) of the GH2 exo-beta-mannanase from Xanthomonas axonopodis pv. citri [Xanthomonas citri pv. citri str. 306] |
| 6BYG_A | 1.12e-50 | 16 | 607 | 28 | 677 | Crystalstructure of the nucleophile mutant (E575A) of the GH2 exo-beta-mannanase from Xanthomonas axonopodis pv. citri [Xanthomonas citri pv. citri str. 306],6BYG_B Crystal structure of the nucleophile mutant (E575A) of the GH2 exo-beta-mannanase from Xanthomonas axonopodis pv. citri [Xanthomonas citri pv. citri str. 306] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| O00462 | 2.72e-47 | 11 | 582 | 32 | 671 | Beta-mannosidase OS=Homo sapiens OX=9606 GN=MANBA PE=1 SV=3 |
| Q4FZV0 | 9.02e-47 | 12 | 518 | 31 | 596 | Beta-mannosidase OS=Rattus norvegicus OX=10116 GN=Manba PE=2 SV=1 |
| Q95327 | 2.93e-46 | 12 | 605 | 31 | 693 | Beta-mannosidase OS=Capra hircus OX=9925 GN=MANBA PE=1 SV=1 |
| Q29444 | 2.32e-45 | 12 | 605 | 31 | 693 | Beta-mannosidase OS=Bos taurus OX=9913 GN=MANBA PE=1 SV=1 |
| Q8K2I4 | 4.19e-45 | 12 | 518 | 31 | 596 | Beta-mannosidase OS=Mus musculus OX=10090 GN=Manba PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
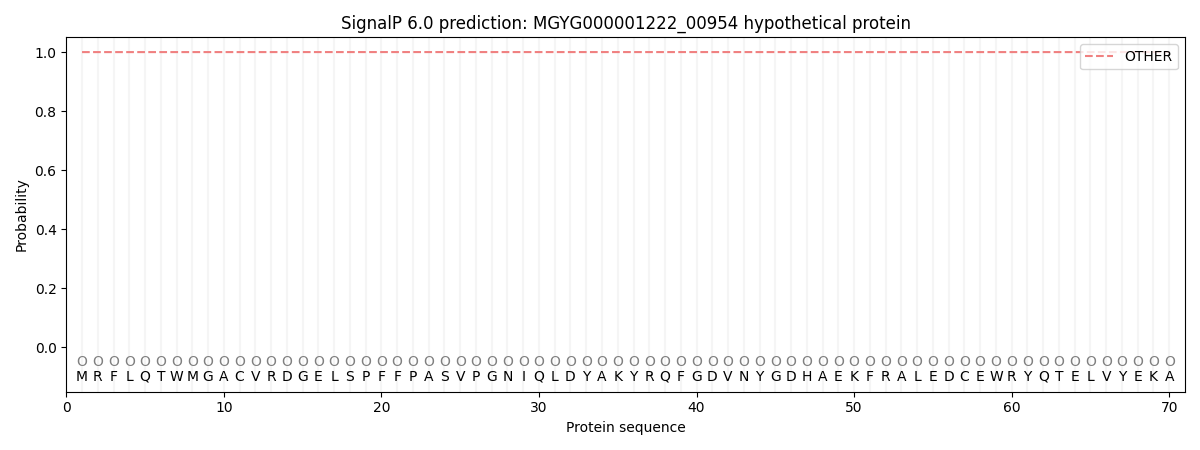
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000043 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
