You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001261_03184
You are here: Home > Sequence: MGYG000001261_03184
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Caulobacter sp903900155 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Proteobacteria; Alphaproteobacteria; Caulobacterales; Caulobacteraceae; Caulobacter; Caulobacter sp903900155 | |||||||||||
| CAZyme ID | MGYG000001261_03184 | |||||||||||
| CAZy Family | GH5 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 1832; End: 3520 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH5 | 82 | 347 | 3.2e-105 | 0.980544747081712 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam00150 | Cellulase | 7.90e-30 | 94 | 342 | 25 | 266 | Cellulase (glycosyl hydrolase family 5). |
| COG2730 | BglC | 1.90e-17 | 87 | 294 | 68 | 250 | Aryl-phospho-beta-D-glucosidase BglC, GH1 family [Carbohydrate transport and metabolism]. |
| cd04080 | CBM6_cellulase-like | 2.81e-15 | 431 | 550 | 4 | 134 | Carbohydrate Binding Module 6 (CBM6); appended to glycoside hydrolase (GH) domains, including GH5 (cellulase). This family includes carbohydrate binding module 6 (CBM6) domains that are appended to several glycoside hydrolase (GH) domains, including GH5 (cellulase) and GH16, as well as to coagulation factor 5/8 carbohydrate-binding domains. CBM6s are non-catalytic carbohydrate binding domains that facilitate the strong binding of the GH catalytic modules with their dedicated, insoluble substrates. The CBM6s are appended to GHs that display a diversity of substrate specificities. For some members of this family information is available about the specific substrates of the appended GH domains. It includes the CBM domains of various enzymes involved in cell wall degradation including, an extracellular beta-1,3-glucanase from Lysobacter enzymogenes encoded by the gluC gene (its catalytic domain belongs to the GH16 family), the tandem CBM domains of Pseudomonas sp. PE2 beta-1,3(4)-glucanase A (its catalytic domain also belongs to GH16), and a family 6 CBM from Cellvibrio mixtus Endoglucanase 5A (CmCBM6) which binds to the beta1,4-beta1,3-mixed linked glucans lichenan, and barley beta-glucan, cello-oligosaccharides, insoluble forms of cellulose, the beta1,3-glucan laminarin, and xylooligosaccharides, and the CBM6 of Fibrobacter succinogenes S85 XynD xylanase, appended to a GH10 domain, and Cellvibrio japonicas Cel5G appended to a GH5 (cellulase) domain. GH5 (cellulase) family includes enzymes with several known activities such as endoglucanase, beta-mannanase, and xylanase, which are involved in the degradation of cellulose and xylans. GH16 family includes enzymes with lichenase, xyloglucan endotransglycosylase (XET), and beta-agarase activities. CBM6 is an unusual CBM as it represents a chimera of two distinct binding sites with different modes of binding: binding site I within the loop regions and binding site II on the concave face of the beta-sandwich fold. For CmCBM6 it has been shown that these two binding sites have different ligand specificities. |
| cd04082 | CBM35_pectate_lyase-like | 0.006 | 489 | 551 | 30 | 114 | Carbohydrate Binding Module family 35 (CBM35), pectate lyase-like; appended mainly to enzymes that bind mannan (Man), xylan, glucuronic acid (GlcA) and possibly glucans. This family includes carbohydrate binding module family 35 (CBM35) domains that are non-catalytic carbohydrate binding domains that are appended mainly to enzymes that bind mannan (Man), xylan, glucuronic acid (GlcA) and possibly glucans. Included in this family are CBM35s of pectate lyases, including pectate lyase 10A from Cellvibrio japonicas, these enzymes release delta-4,5-anhydrogalaturonic acid (delta4,5-GalA) from pectin, thus identifying a signature molecule for plant cell wall degradation. CBM35s are unique in that they display conserved specificity through extensive sequence similarity but divergent function through their appended catalytic modules. They are known to bind alpha-D-galactose (Gal), mannan (Man), xylan, glucuronic acid (GlcA), a beta-polymer of mannose, and possibly glucans, forming four subfamilies based on general ligand specificities (galacto, urono, manno, and gluco configurations). In contrast to most CBMs that are generally rigid proteins, CBM35 undergoes significant conformational change upon ligand binding. Some CBM35s bind their ligands in a calcium-dependent manner, especially those binding uronic acids. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| SBV32487.1 | 7.33e-234 | 5 | 511 | 6 | 510 |
| AJA09151.1 | 1.20e-232 | 1 | 511 | 1 | 510 |
| APZ97684.1 | 4.85e-232 | 9 | 512 | 7 | 511 |
| QNO27624.1 | 1.43e-231 | 8 | 512 | 4 | 511 |
| QCB54589.1 | 1.99e-226 | 1 | 511 | 1 | 510 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1CEC_A | 4.81e-21 | 80 | 350 | 15 | 321 | ChainA, ENDOGLUCANASE CELC [Acetivibrio thermocellus] |
| 1CEN_A | 1.18e-20 | 80 | 350 | 15 | 321 | ChainA, CELLULASE CELC [Acetivibrio thermocellus],1CEO_A Chain A, CELLULASE CELC [Acetivibrio thermocellus] |
| 6JYZ_A | 5.21e-16 | 14 | 342 | 14 | 360 | ChainA, Putative secreted endoglycosylceramidase [Rhodococcus hoagii 103S] |
| 3AMC_A | 1.38e-13 | 100 | 342 | 40 | 286 | Crystalstructures of Thermotoga maritima Cel5A, apo form and dimer/au [Thermotoga maritima MSB8],3AMC_B Crystal structures of Thermotoga maritima Cel5A, apo form and dimer/au [Thermotoga maritima MSB8],3AMD_A Crystal structures of Thermotoga maritima Cel5A, apo form and tetramer/au [Thermotoga maritima MSB8],3AMD_B Crystal structures of Thermotoga maritima Cel5A, apo form and tetramer/au [Thermotoga maritima MSB8],3AMD_C Crystal structures of Thermotoga maritima Cel5A, apo form and tetramer/au [Thermotoga maritima MSB8],3AMD_D Crystal structures of Thermotoga maritima Cel5A, apo form and tetramer/au [Thermotoga maritima MSB8],3MMU_A Crystal structure of endoglucanase Cel5A from the hyperthermophilic Thermotoga maritima [Thermotoga maritima],3MMU_B Crystal structure of endoglucanase Cel5A from the hyperthermophilic Thermotoga maritima [Thermotoga maritima],3MMU_C Crystal structure of endoglucanase Cel5A from the hyperthermophilic Thermotoga maritima [Thermotoga maritima],3MMU_D Crystal structure of endoglucanase Cel5A from the hyperthermophilic Thermotoga maritima [Thermotoga maritima],3MMU_E Crystal structure of endoglucanase Cel5A from the hyperthermophilic Thermotoga maritima [Thermotoga maritima],3MMU_F Crystal structure of endoglucanase Cel5A from the hyperthermophilic Thermotoga maritima [Thermotoga maritima],3MMU_G Crystal structure of endoglucanase Cel5A from the hyperthermophilic Thermotoga maritima [Thermotoga maritima],3MMU_H Crystal structure of endoglucanase Cel5A from the hyperthermophilic Thermotoga maritima [Thermotoga maritima],3MMW_A Crystal structure of endoglucanase Cel5A from the hyperthermophilic Thermotoga maritima [Thermotoga maritima],3MMW_B Crystal structure of endoglucanase Cel5A from the hyperthermophilic Thermotoga maritima [Thermotoga maritima],3MMW_C Crystal structure of endoglucanase Cel5A from the hyperthermophilic Thermotoga maritima [Thermotoga maritima],3MMW_D Crystal structure of endoglucanase Cel5A from the hyperthermophilic Thermotoga maritima [Thermotoga maritima] |
| 3AMG_A | 8.02e-13 | 100 | 342 | 40 | 286 | Crystalstructures of Thermotoga maritima Cel5A in complex with Cellobiose substrate, mutant form [Thermotoga maritima MSB8],3AMG_B Crystal structures of Thermotoga maritima Cel5A in complex with Cellobiose substrate, mutant form [Thermotoga maritima MSB8] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| W8QRE4 | 5.42e-42 | 24 | 402 | 3 | 404 | Beta-xylosidase OS=Phanerodontia chrysosporium OX=2822231 GN=Xyl5 PE=1 SV=2 |
| P23340 | 1.07e-20 | 80 | 350 | 15 | 321 | Endoglucanase C307 OS=Clostridium sp. (strain F1) OX=1508 GN=celC307 PE=1 SV=1 |
| A3DJ77 | 1.07e-20 | 80 | 350 | 15 | 321 | Endoglucanase C OS=Acetivibrio thermocellus (strain ATCC 27405 / DSM 1237 / JCM 9322 / NBRC 103400 / NCIMB 10682 / NRRL B-4536 / VPI 7372) OX=203119 GN=celC PE=3 SV=1 |
| P0C2S3 | 4.79e-20 | 80 | 350 | 15 | 321 | Endoglucanase C OS=Acetivibrio thermocellus OX=1515 GN=celC PE=1 SV=1 |
| A2RAR6 | 1.11e-18 | 11 | 303 | 12 | 287 | Probable glucan 1,3-beta-glucosidase A OS=Aspergillus niger (strain CBS 513.88 / FGSC A1513) OX=425011 GN=exgA PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
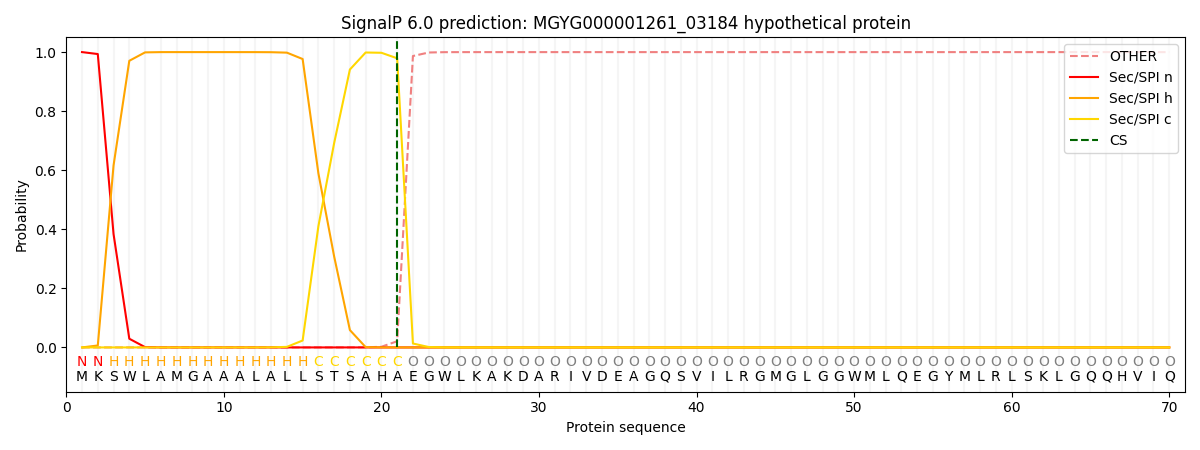
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000312 | 0.998933 | 0.000191 | 0.000189 | 0.000181 | 0.000161 |
