You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001262_01543
You are here: Home > Sequence: MGYG000001262_01543
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Sphingomonas ginsenosidimutans | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Proteobacteria; Alphaproteobacteria; Sphingomonadales; Sphingomonadaceae; Sphingomonas; Sphingomonas ginsenosidimutans | |||||||||||
| CAZyme ID | MGYG000001262_01543 | |||||||||||
| CAZy Family | CBM48 | |||||||||||
| CAZyme Description | 1,4-alpha-glucan branching enzyme GlgB | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 11199; End: 13367 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH13 | 273 | 577 | 3.1e-155 | 0.9966777408637874 |
| CBM48 | 119 | 205 | 1.3e-20 | 0.8947368421052632 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| PRK05402 | PRK05402 | 0.0 | 3 | 722 | 1 | 726 | 1,4-alpha-glucan branching protein GlgB. |
| PRK14705 | PRK14705 | 0.0 | 90 | 722 | 593 | 1223 | glycogen branching enzyme; Provisional |
| PRK12313 | PRK12313 | 0.0 | 104 | 722 | 11 | 630 | 1,4-alpha-glucan branching protein GlgB. |
| cd11322 | AmyAc_Glg_BE | 0.0 | 208 | 613 | 1 | 402 | Alpha amylase catalytic domain found in the Glycogen branching enzyme (also called 1,4-alpha-glucan branching enzyme). The glycogen branching enzyme catalyzes the third step of glycogen biosynthesis by the cleavage of an alpha-(1,4)-glucosidic linkage and the formation a new alpha-(1,6)-branch by subsequent transfer of cleaved oligosaccharide. They are part of a group called branching enzymes which catalyze the formation of alpha-1,6 branch points in either glycogen or starch. This group includes proteins from bacteria, eukaryotes, and archaea. The Alpha-amylase family comprises the largest family of glycoside hydrolases (GH), with the majority of enzymes acting on starch, glycogen, and related oligo- and polysaccharides. These proteins catalyze the transformation of alpha-1,4 and alpha-1,6 glucosidic linkages with retention of the anomeric center. The protein is described as having 3 domains: A, B, C. A is a (beta/alpha) 8-barrel; B is a loop between the beta 3 strand and alpha 3 helix of A; C is the C-terminal extension characterized by a Greek key. The majority of the enzymes have an active site cleft found between domains A and B where a triad of catalytic residues (Asp, Glu and Asp) performs catalysis. Other members of this family have lost the catalytic activity as in the case of the human 4F2hc, or only have 2 residues that serve as the catalytic nucleophile and the acid/base, such as Thermus A4 beta-galactosidase with 2 Glu residues (GH42) and human alpha-galactosidase with 2 Asp residues (GH31). The family members are quite extensive and include: alpha amylase, maltosyltransferase, cyclodextrin glycotransferase, maltogenic amylase, neopullulanase, isoamylase, 1,4-alpha-D-glucan maltotetrahydrolase, 4-alpha-glucotransferase, oligo-1,6-glucosidase, amylosucrase, sucrose phosphorylase, and amylomaltase. |
| COG0296 | GlgB | 0.0 | 95 | 717 | 1 | 626 | 1,4-alpha-glucan branching enzyme [Carbohydrate transport and metabolism]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| AIT05614.1 | 0.0 | 1 | 720 | 1 | 720 |
| ANC88017.1 | 0.0 | 1 | 720 | 1 | 720 |
| AOW23523.1 | 0.0 | 1 | 720 | 1 | 720 |
| ATI54467.1 | 0.0 | 1 | 720 | 1 | 720 |
| BCA62629.1 | 0.0 | 1 | 720 | 1 | 719 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1M7X_A | 9.15e-249 | 107 | 720 | 1 | 615 | TheX-ray Crystallographic Structure of Branching Enzyme [Escherichia coli],1M7X_B The X-ray Crystallographic Structure of Branching Enzyme [Escherichia coli],1M7X_C The X-ray Crystallographic Structure of Branching Enzyme [Escherichia coli],1M7X_D The X-ray Crystallographic Structure of Branching Enzyme [Escherichia coli] |
| 4LPC_A | 2.03e-246 | 112 | 720 | 1 | 610 | CrystalStructure of E.Coli Branching Enzyme in complex with maltoheptaose [Escherichia coli],4LPC_B Crystal Structure of E.Coli Branching Enzyme in complex with maltoheptaose [Escherichia coli],4LPC_C Crystal Structure of E.Coli Branching Enzyme in complex with maltoheptaose [Escherichia coli],4LPC_D Crystal Structure of E.Coli Branching Enzyme in complex with maltoheptaose [Escherichia coli],4LQ1_A Crystal Structure of E.Coli Branching Enzyme in complex with maltohexaose [Escherichia coli],4LQ1_B Crystal Structure of E.Coli Branching Enzyme in complex with maltohexaose [Escherichia coli],4LQ1_C Crystal Structure of E.Coli Branching Enzyme in complex with maltohexaose [Escherichia coli],4LQ1_D Crystal Structure of E.Coli Branching Enzyme in complex with maltohexaose [Escherichia coli],5E6Y_A Crystal structure of E.Coli branching enzyme in complex with alpha cyclodextrin [Escherichia coli O139:H28 str. E24377A],5E6Y_B Crystal structure of E.Coli branching enzyme in complex with alpha cyclodextrin [Escherichia coli O139:H28 str. E24377A],5E6Y_C Crystal structure of E.Coli branching enzyme in complex with alpha cyclodextrin [Escherichia coli O139:H28 str. E24377A],5E6Y_D Crystal structure of E.Coli branching enzyme in complex with alpha cyclodextrin [Escherichia coli O139:H28 str. E24377A],5E6Z_A Crystal structure of Ecoli Branching Enzyme with beta cyclodextrin [Escherichia coli O139:H28 str. E24377A],5E6Z_B Crystal structure of Ecoli Branching Enzyme with beta cyclodextrin [Escherichia coli O139:H28 str. E24377A],5E6Z_C Crystal structure of Ecoli Branching Enzyme with beta cyclodextrin [Escherichia coli O139:H28 str. E24377A],5E6Z_D Crystal structure of Ecoli Branching Enzyme with beta cyclodextrin [Escherichia coli O139:H28 str. E24377A],5E70_A Crystal structure of Ecoli Branching Enzyme with gamma cyclodextrin [Escherichia coli O139:H28 str. E24377A],5E70_B Crystal structure of Ecoli Branching Enzyme with gamma cyclodextrin [Escherichia coli O139:H28 str. E24377A],5E70_C Crystal structure of Ecoli Branching Enzyme with gamma cyclodextrin [Escherichia coli O139:H28 str. E24377A],5E70_D Crystal structure of Ecoli Branching Enzyme with gamma cyclodextrin [Escherichia coli O139:H28 str. E24377A] |
| 3K1D_A | 1.34e-231 | 1 | 706 | 3 | 701 | Crystalstructure of glycogen branching enzyme synonym: 1,4-alpha-D-glucan:1,4-alpha-D-GLUCAN 6-glucosyl-transferase from mycobacterium tuberculosis H37RV [Mycobacterium tuberculosis H37Rv] |
| 5GQW_A | 6.52e-229 | 5 | 721 | 27 | 775 | Crystalstructure of branching enzyme W610N mutant from Cyanothece sp. ATCC 51142 [Crocosphaera subtropica ATCC 51142],5GQX_A Crystal structure of branching enzyme W610N mutant from Cyanothece sp. ATCC 51142 in complex with maltoheptaose [Crocosphaera subtropica ATCC 51142] |
| 5GR5_A | 9.23e-229 | 5 | 721 | 27 | 775 | Crystalstructure of branching enzyme W610A mutant from Cyanothece sp. ATCC 51142 [Crocosphaera subtropica ATCC 51142] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P52979 | 0.0 | 6 | 717 | 19 | 730 | 1,4-alpha-glucan branching enzyme GlgB OS=Rhizobium radiobacter OX=358 GN=glgB PE=3 SV=2 |
| Q1MBS9 | 0.0 | 5 | 717 | 18 | 731 | 1,4-alpha-glucan branching enzyme GlgB 1 OS=Rhizobium leguminosarum bv. viciae (strain 3841) OX=216596 GN=glgB1 PE=3 SV=1 |
| Q8U8L4 | 0.0 | 6 | 717 | 19 | 731 | 1,4-alpha-glucan branching enzyme GlgB OS=Agrobacterium fabrum (strain C58 / ATCC 33970) OX=176299 GN=glgB PE=3 SV=1 |
| Q2G7S5 | 0.0 | 1 | 720 | 1 | 720 | 1,4-alpha-glucan branching enzyme GlgB OS=Novosphingobium aromaticivorans (strain ATCC 700278 / DSM 12444 / CCUG 56034 / CIP 105152 / NBRC 16084 / F199) OX=279238 GN=glgB PE=3 SV=1 |
| Q92M14 | 0.0 | 4 | 717 | 18 | 733 | 1,4-alpha-glucan branching enzyme GlgB OS=Rhizobium meliloti (strain 1021) OX=266834 GN=glgB PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
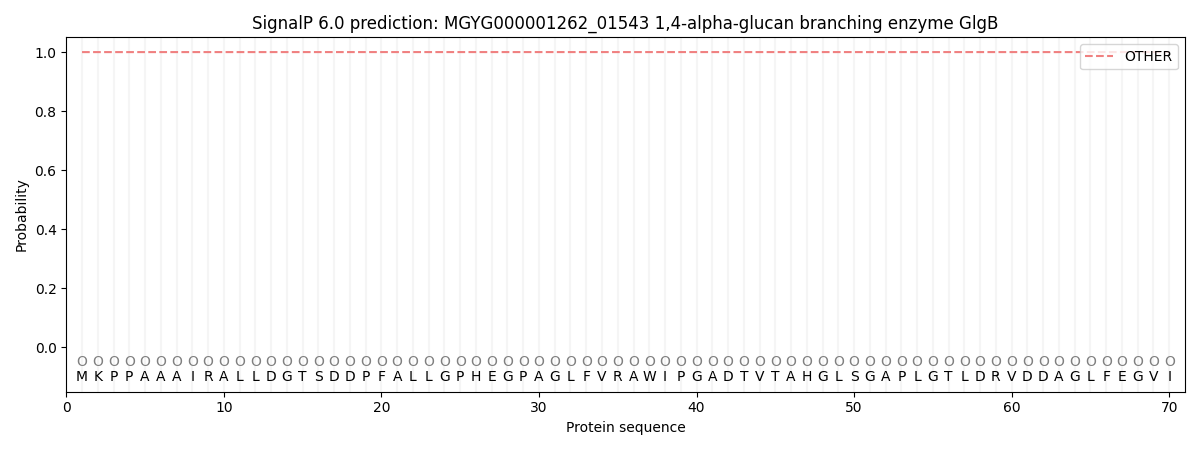
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000026 | 0.000039 | 0.000001 | 0.000000 | 0.000000 | 0.000000 |
