You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001265_02648
You are here: Home > Sequence: MGYG000001265_02648
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Methylobacterium sp002778835 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Proteobacteria; Alphaproteobacteria; Rhizobiales; Beijerinckiaceae; Methylobacterium; Methylobacterium sp002778835 | |||||||||||
| CAZyme ID | MGYG000001265_02648 | |||||||||||
| CAZy Family | GT19 | |||||||||||
| CAZyme Description | Lipid-A-disaccharide synthase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 50; End: 673 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GT19 | 2 | 168 | 8.1e-47 | 0.4576271186440678 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| PRK00025 | lpxB | 4.63e-41 | 2 | 195 | 191 | 380 | lipid-A-disaccharide synthase; Reviewed |
| COG0763 | LpxB | 1.48e-39 | 2 | 192 | 193 | 381 | Lipid A disaccharide synthetase [Cell wall/membrane/envelope biogenesis]. |
| pfam02684 | LpxB | 1.56e-26 | 2 | 172 | 190 | 361 | Lipid-A-disaccharide synthetase. This is a family of lipid-A-disaccharide synthetases, EC:2.4.2.128. These enzymes catalyze the reaction: UDP-2,3-bis(3-hydroxytetradecanoyl) glucosamine + 2,3-bis(3-hydroxytetradecanoyl)-beta-D-glucosaminyl 1-phosphate <=> UDP + 2,3-bis(3-hydroxytetradecanoyl)-D-glucosaminyl-1,6 -beta-D-2,3-bis(3-hydroxytetradecanoyl)-beta-D-glucosaminyl 1-phosphate. These enzymes catalyze the fist disaccharide step in the synthesis of lipid-A-disaccharide. |
| PRK14089 | PRK14089 | 2.91e-16 | 3 | 140 | 173 | 307 | lipid-A-disaccharide synthase. |
| PRK01021 | lpxB | 0.003 | 81 | 140 | 497 | 560 | lipid-A-disaccharide synthase; Reviewed |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| AWI91282.1 | 5.24e-86 | 2 | 191 | 195 | 384 |
| QIJ77275.1 | 7.42e-86 | 2 | 191 | 195 | 384 |
| QIJ82179.1 | 7.42e-86 | 2 | 191 | 195 | 384 |
| ACS42731.1 | 1.49e-85 | 2 | 191 | 195 | 384 |
| ACB83314.1 | 2.11e-85 | 2 | 191 | 195 | 384 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5W8N_A | 2.08e-21 | 2 | 191 | 196 | 382 | LipidA Disaccharide Synthase (LpxB)-6 solubilizing mutations [Escherichia coli BL21(DE3)] |
| 5W8S_A | 2.08e-21 | 2 | 191 | 196 | 382 | LipidA Disaccharide Synthase (LpxB)-7 solubilizing mutations [Escherichia coli BL21(DE3)],5W8X_A Lipid A Disaccharide Synthase (LpxB)-7 solubilizing mutations-Bound to UDP [Escherichia coli BL21(DE3)] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q2IW93 | 1.06e-44 | 1 | 192 | 202 | 392 | Lipid-A-disaccharide synthase OS=Rhodopseudomonas palustris (strain HaA2) OX=316058 GN=lpxB PE=3 SV=1 |
| Q3SRI5 | 9.29e-43 | 1 | 192 | 201 | 391 | Lipid-A-disaccharide synthase OS=Nitrobacter winogradskyi (strain ATCC 25391 / DSM 10237 / CIP 104748 / NCIMB 11846 / Nb-255) OX=323098 GN=lpxB PE=3 SV=1 |
| Q6N5R2 | 1.31e-41 | 1 | 191 | 202 | 391 | Lipid-A-disaccharide synthase OS=Rhodopseudomonas palustris (strain ATCC BAA-98 / CGA009) OX=258594 GN=lpxB PE=3 SV=1 |
| Q1QMM4 | 2.74e-41 | 1 | 188 | 201 | 387 | Lipid-A-disaccharide synthase OS=Nitrobacter hamburgensis (strain DSM 10229 / NCIMB 13809 / X14) OX=323097 GN=lpxB PE=3 SV=1 |
| Q89KQ7 | 4.99e-41 | 1 | 192 | 199 | 389 | Lipid-A-disaccharide synthase OS=Bradyrhizobium diazoefficiens (strain JCM 10833 / BCRC 13528 / IAM 13628 / NBRC 14792 / USDA 110) OX=224911 GN=lpxB PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
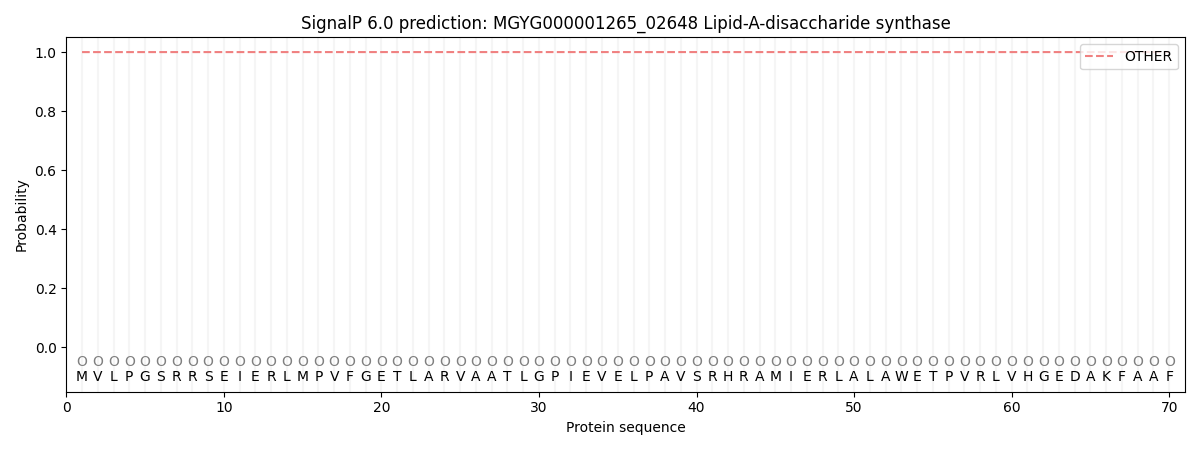
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000055 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
