You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001279_02964
You are here: Home > Sequence: MGYG000001279_02964
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Pseudomonas_E fulva | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Proteobacteria; Gammaproteobacteria; Pseudomonadales; Pseudomonadaceae; Pseudomonas_E; Pseudomonas_E fulva | |||||||||||
| CAZyme ID | MGYG000001279_02964 | |||||||||||
| CAZy Family | GH23 | |||||||||||
| CAZyme Description | Membrane-bound lytic murein transglycosylase D | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 2; End: 1432 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH23 | 116 | 260 | 1.3e-24 | 0.9333333333333333 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| PRK10783 | mltD | 4.10e-131 | 29 | 474 | 13 | 448 | membrane-bound lytic murein transglycosylase D; Provisional |
| cd16894 | MltD-like | 5.63e-65 | 124 | 254 | 1 | 129 | Membrane-bound lytic murein transglycosylase D and similar proteins. Lytic transglycosylases (LT) catalyze the cleavage of the beta-1,4-glycosidic bond between N-acetylmuramic acid (MurNAc) and N-acetyl-D-glucosamine (GlcNAc). Membrane-bound lytic murein transglycosylase D protein (MltD) family members may have one or more small LysM domains, which may contribute to peptidoglycan binding. Unlike the similar "goose-type" lysozymes, LTs also make a new glycosidic bond with the C6 hydroxyl group of the same muramic acid residue. Proteins similar to this family include the soluble and insoluble membrane-bound LTs in bacteria, the LTs in bacteriophage lambda, as well as the eukaryotic "goose-type" lysozymes (goose egg-white lysozyme; GEWL). |
| pfam01464 | SLT | 1.02e-29 | 129 | 220 | 11 | 102 | Transglycosylase SLT domain. This family is distantly related to pfam00062. Members are found in phages, type II, type III and type IV secretion systems. |
| cd13401 | Slt70-like | 3.05e-25 | 131 | 251 | 19 | 144 | 70kDa soluble lytic transglycosylase (Slt70) and similar proteins. Catalytic domain of the 70kda soluble lytic transglycosylase (LT)-like proteins, which also have an N-terminal U-shaped U-domain and a linker L-domain. LTs catalyze the cleavage of the beta-1,4-glycosidic bond between N-acetylmuramic acid (MurNAc) and N-acetyl-D-glucosamine (GlcNAc), as do "goose-type" lysozymes. However, in addition to this, they also make a new glycosidic bond with the C6 hydroxyl group of the same muramic acid residue. Proteins similar to this family include the soluble and insoluble membrane-bound LTs in bacteria and the LTs in bacteriophage lambda. |
| cd00254 | LT-like | 6.77e-23 | 134 | 254 | 2 | 111 | lytic transglycosylase(LT)-like domain. Members include the soluble and insoluble membrane-bound LTs in bacteria and LTs in bacteriophage lambda. LTs catalyze the cleavage of the beta-1,4-glycosidic bond between N-acetylmuramic acid (MurNAc) and N-acetyl-D-glucosamine (GlcNAc), as do "goose-type" lysozymes. However, in addition to this, they also make a new glycosidic bond with the C6 hydroxyl group of the same muramic acid residue. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QPH47608.1 | 0.0 | 1 | 476 | 1 | 476 |
| QDC03963.1 | 0.0 | 1 | 476 | 1 | 476 |
| QPH42545.1 | 0.0 | 1 | 476 | 1 | 476 |
| CRN07691.1 | 0.0 | 1 | 476 | 1 | 476 |
| AVF56107.1 | 0.0 | 1 | 476 | 1 | 476 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5OHU_A | 5.64e-08 | 140 | 276 | 503 | 640 | TheX-ray Structure of Lytic Transglycosylase Slt from Pseudomonas aeruginosa [Pseudomonas aeruginosa] |
| 6FBT_A | 9.59e-08 | 140 | 255 | 474 | 591 | ChainA, Lytic murein transglycosylase [Pseudomonas aeruginosa] |
| 6FC4_A | 1.27e-07 | 140 | 276 | 475 | 612 | ChainA, Soluble lytic murein transglycosylase [Pseudomonas aeruginosa] |
| 6FCQ_A | 1.27e-07 | 140 | 276 | 474 | 611 | ChainA, Soluble lytic murein transglycosylase [Pseudomonas aeruginosa],6FCR_A Chain A, Soluble lytic murein transglycosylase [Pseudomonas aeruginosa],6FCS_A Chain A, Soluble lytic murein transglycosylase [Pseudomonas aeruginosa],6FCU_A Chain A, Soluble lytic murein transglycosylase [Pseudomonas aeruginosa] |
| 7EYB_I | 2.54e-06 | 140 | 268 | 37 | 162 | ChainI, Peptidoglycan transglycosylase gp16 [Escherichia phage T7],7EYB_J Chain J, Peptidoglycan transglycosylase gp16 [Escherichia phage T7],7EYB_K Chain K, Peptidoglycan transglycosylase gp16 [Escherichia phage T7],7EYB_L Chain L, Peptidoglycan transglycosylase gp16 [Escherichia phage T7],7K5C_B Chain B, Peptidoglycan transglycosylase gp16 [Escherichia phage T7],7K5C_D Chain D, Peptidoglycan transglycosylase gp16 [Escherichia phage T7],7K5C_F Chain F, Peptidoglycan transglycosylase gp16 [Escherichia phage T7],7K5C_G Chain G, Peptidoglycan transglycosylase gp16 [Escherichia phage T7],7K5C_J Chain J, Peptidoglycan transglycosylase gp16 [Escherichia phage T7],7K5C_L Chain L, Peptidoglycan transglycosylase gp16 [Escherichia phage T7] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P0AEZ7 | 5.53e-87 | 29 | 474 | 13 | 444 | Membrane-bound lytic murein transglycosylase D OS=Escherichia coli (strain K12) OX=83333 GN=mltD PE=1 SV=1 |
| P0AEZ8 | 5.53e-87 | 29 | 474 | 13 | 444 | Membrane-bound lytic murein transglycosylase D OS=Escherichia coli O6:H1 (strain CFT073 / ATCC 700928 / UPEC) OX=199310 GN=mltD PE=3 SV=1 |
| P32820 | 1.84e-30 | 104 | 292 | 15 | 203 | Putative tributyltin chloride resistance protein OS=Alteromonas sp. (strain M-1) OX=29457 GN=tbtA PE=3 SV=1 |
| P37710 | 9.49e-10 | 367 | 475 | 567 | 677 | Autolysin OS=Enterococcus faecalis (strain ATCC 700802 / V583) OX=226185 GN=EF_0799 PE=1 SV=2 |
| O31608 | 3.28e-09 | 140 | 252 | 85 | 178 | Putative murein lytic transglycosylase YjbJ OS=Bacillus subtilis (strain 168) OX=224308 GN=yjbJ PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as LIPO
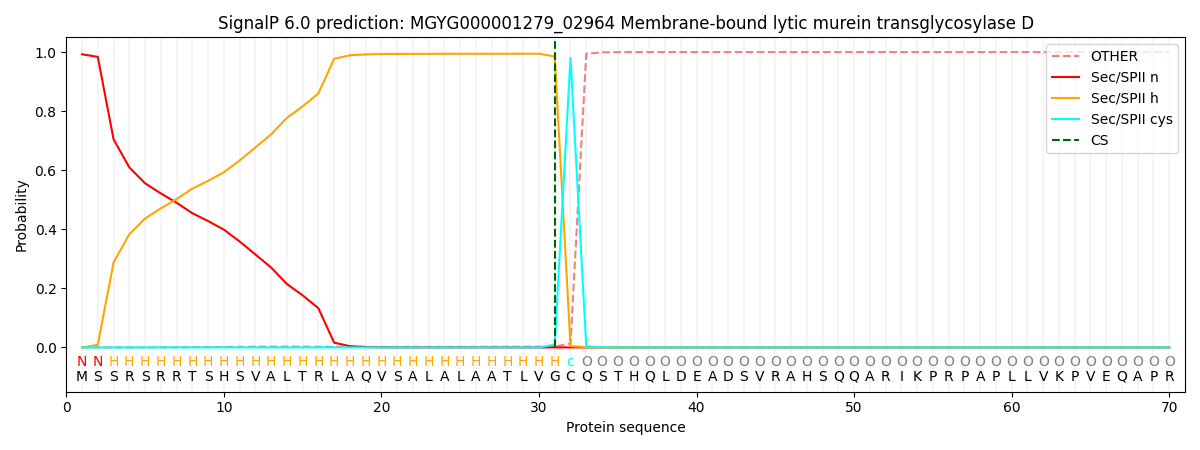
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000024 | 0.003418 | 0.996572 | 0.000004 | 0.000006 | 0.000002 |
