You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001303_01456
You are here: Home > Sequence: MGYG000001303_01456
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
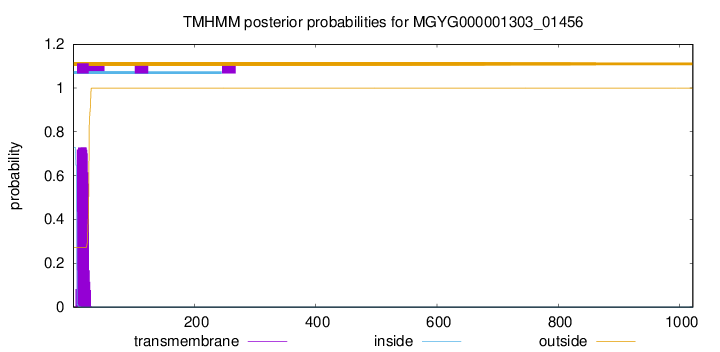
TMHMM annotations
Basic Information help
| Species | Clostridium_AP scindens | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Lachnospirales; Lachnospiraceae; Clostridium_AP; Clostridium_AP scindens | |||||||||||
| CAZyme ID | MGYG000001303_01456 | |||||||||||
| CAZy Family | CBM32 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 3225; End: 6293 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| CBM32 | 178 | 298 | 8.9e-21 | 0.9274193548387096 |
| CBM32 | 472 | 570 | 3.9e-16 | 0.8064516129032258 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam00754 | F5_F8_type_C | 4.43e-19 | 178 | 297 | 4 | 127 | F5/8 type C domain. This domain is also known as the discoidin (DS) domain family. |
| pfam02585 | PIG-L | 1.23e-12 | 597 | 669 | 1 | 79 | GlcNAc-PI de-N-acetylase. Members of this family are related to PIG-L an N-acetylglucosaminylphosphatidylinositol de-N-acetylase (EC:3.5.1.89) that catalyzes the second step in GPI biosynthesis. |
| pfam00754 | F5_F8_type_C | 1.26e-12 | 469 | 566 | 1 | 110 | F5/8 type C domain. This domain is also known as the discoidin (DS) domain family. |
| pfam00754 | F5_F8_type_C | 1.90e-12 | 310 | 428 | 1 | 114 | F5/8 type C domain. This domain is also known as the discoidin (DS) domain family. |
| COG2120 | LmbE | 6.86e-09 | 592 | 669 | 10 | 89 | N-acetylglucosaminyl deacetylase, LmbE family [Carbohydrate transport and metabolism]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QRO38183.1 | 0.0 | 1 | 1022 | 1 | 1022 |
| QBF74995.1 | 0.0 | 1 | 1022 | 1 | 1022 |
| QMW76217.1 | 3.54e-51 | 39 | 506 | 561 | 1091 |
| QIB55915.1 | 3.54e-51 | 39 | 506 | 561 | 1091 |
| QRO38181.1 | 1.88e-50 | 39 | 481 | 567 | 1075 |
Swiss-Prot Hits help
SignalP and Lipop Annotations help
This protein is predicted as SP
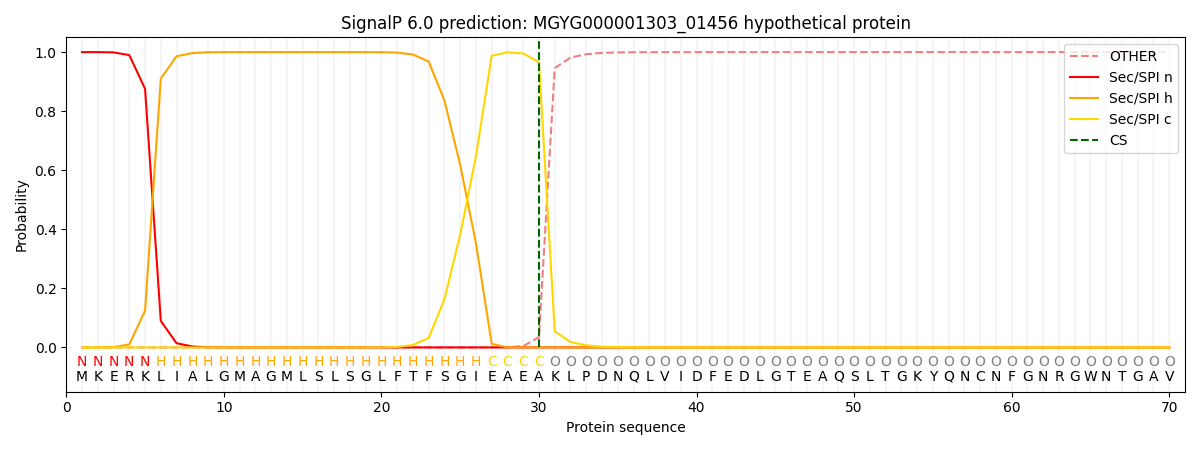
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000364 | 0.998845 | 0.000194 | 0.000222 | 0.000183 | 0.000161 |