You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001306_03108
You are here: Home > Sequence: MGYG000001306_03108
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Phocaeicola coprocola | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Bacteroidaceae; Phocaeicola; Phocaeicola coprocola | |||||||||||
| CAZyme ID | MGYG000001306_03108 | |||||||||||
| CAZy Family | CE1 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 14726; End: 15499 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| CE1 | 34 | 237 | 1.4e-26 | 0.8942731277533039 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG4099 | COG4099 | 1.24e-48 | 16 | 255 | 153 | 386 | Predicted peptidase [General function prediction only]. |
| COG1506 | DAP2 | 7.30e-21 | 23 | 254 | 364 | 616 | Dipeptidyl aminopeptidase/acylaminoacyl peptidase [Amino acid transport and metabolism]. |
| COG0400 | YpfH | 1.07e-14 | 54 | 232 | 19 | 186 | Predicted esterase [General function prediction only]. |
| pfam02230 | Abhydrolase_2 | 2.45e-12 | 44 | 239 | 4 | 203 | Phospholipase/Carboxylesterase. This family consists of both phospholipases and carboxylesterases with broad substrate specificity, and is structurally related to alpha/beta hydrolases pfam00561. |
| pfam00326 | Peptidase_S9 | 7.77e-12 | 136 | 251 | 53 | 206 | Prolyl oligopeptidase family. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| ABS60377.1 | 1.18e-50 | 15 | 257 | 1 | 246 |
| BCI61582.1 | 1.82e-47 | 21 | 255 | 808 | 1042 |
| QDU56037.1 | 5.24e-32 | 51 | 256 | 817 | 1007 |
| VTR91196.1 | 2.65e-31 | 23 | 256 | 32 | 239 |
| ACR12533.1 | 4.30e-31 | 36 | 255 | 63 | 280 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3DOH_A | 2.36e-47 | 33 | 256 | 154 | 380 | CrystalStructure of a Thermostable Esterase [Thermotoga maritima],3DOH_B Crystal Structure of a Thermostable Esterase [Thermotoga maritima],3DOI_A Crystal Structure of a Thermostable Esterase complex with paraoxon [Thermotoga maritima],3DOI_B Crystal Structure of a Thermostable Esterase complex with paraoxon [Thermotoga maritima] |
| 4Q82_A | 2.10e-12 | 51 | 256 | 78 | 277 | CrystalStructure of Phospholipase/Carboxylesterase from Haliangium ochraceum [Haliangium ochraceum DSM 14365],4Q82_B Crystal Structure of Phospholipase/Carboxylesterase from Haliangium ochraceum [Haliangium ochraceum DSM 14365] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q0J968 | 1.11e-07 | 47 | 251 | 3 | 213 | Probable carboxylesterase Os04g0669600 OS=Oryza sativa subsp. japonica OX=39947 GN=Os04g0669600 PE=2 SV=1 |
| P52090 | 2.20e-07 | 130 | 195 | 129 | 192 | Poly(3-hydroxyalkanoate) depolymerase C OS=Paucimonas lemoignei OX=29443 GN=phaZ1 PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
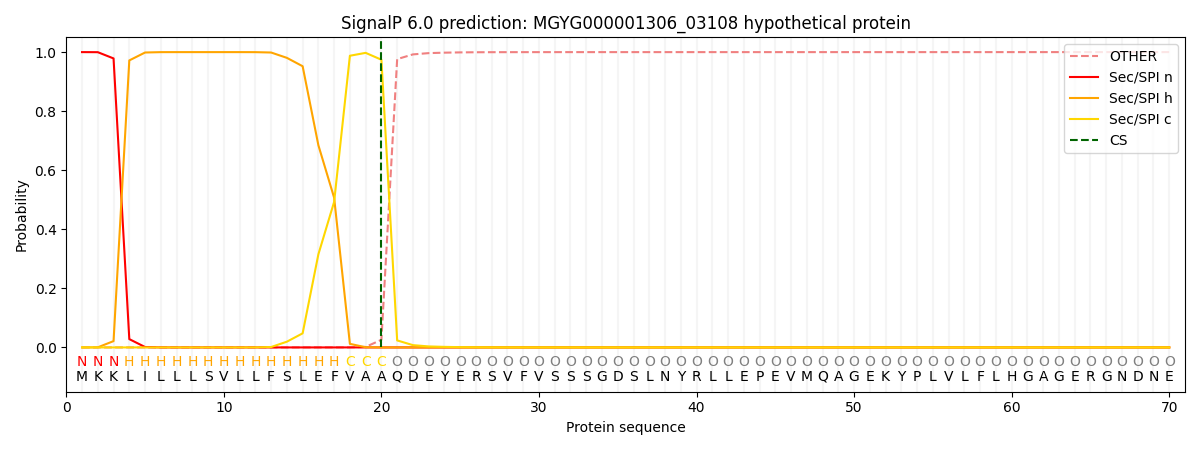
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000290 | 0.999006 | 0.000196 | 0.000157 | 0.000154 | 0.000149 |
