You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001313_02444
You are here: Home > Sequence: MGYG000001313_02444
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Bacteroides eggerthii | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Bacteroidaceae; Bacteroides; Bacteroides eggerthii | |||||||||||
| CAZyme ID | MGYG000001313_02444 | |||||||||||
| CAZy Family | GH10 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 734825; End: 736984 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH10 | 450 | 714 | 4.2e-48 | 0.6864686468646864 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| smart00633 | Glyco_10 | 7.17e-33 | 452 | 712 | 54 | 263 | Glycosyl hydrolase family 10. |
| pfam00331 | Glyco_hydro_10 | 1.43e-31 | 452 | 714 | 96 | 310 | Glycosyl hydrolase family 10. |
| COG3693 | XynA | 9.67e-24 | 450 | 719 | 117 | 344 | Endo-1,4-beta-xylanase, GH35 family [Carbohydrate transport and metabolism]. |
| pfam00331 | Glyco_hydro_10 | 4.90e-13 | 63 | 164 | 11 | 106 | Glycosyl hydrolase family 10. |
| COG3693 | XynA | 1.89e-08 | 87 | 187 | 58 | 175 | Endo-1,4-beta-xylanase, GH35 family [Carbohydrate transport and metabolism]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QRQ48288.1 | 0.0 | 1 | 719 | 1 | 719 |
| QUT46089.1 | 0.0 | 1 | 719 | 1 | 719 |
| ADD61487.1 | 6.72e-309 | 1 | 719 | 1 | 757 |
| ADD61481.1 | 6.72e-309 | 1 | 719 | 1 | 757 |
| QIU93949.1 | 5.04e-295 | 9 | 716 | 2 | 725 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 4MGS_A | 2.85e-21 | 154 | 297 | 1 | 150 | BiXyn10ACBM1 APO [Bacteroides intestinalis DSM 17393] |
| 4QPW_A | 2.49e-17 | 164 | 294 | 5 | 141 | BiXyn10ACBM1 with Xylohexaose Bound [Bacteroides intestinalis DSM 17393] |
| 2F8Q_A | 3.45e-16 | 450 | 709 | 115 | 348 | Analkali thermostable F/10 xylanase from alkalophilic Bacillus sp. NG-27 [Bacillus sp. NG-27],2F8Q_B An alkali thermostable F/10 xylanase from alkalophilic Bacillus sp. NG-27 [Bacillus sp. NG-27] |
| 2FGL_A | 3.49e-16 | 450 | 709 | 116 | 349 | Analkali thermostable F/10 xylanase from alkalophilic Bacillus sp. NG-27 [Bacillus sp. NG-27],2FGL_B An alkali thermostable F/10 xylanase from alkalophilic Bacillus sp. NG-27 [Bacillus sp. NG-27] |
| 4QCE_A | 3.53e-16 | 450 | 709 | 117 | 350 | Crystalstructure of recombinant alkali thermostable GH10 xylanase from Bacillus sp. NG-27 [Bacillus sp. NG-27],4QCE_B Crystal structure of recombinant alkali thermostable GH10 xylanase from Bacillus sp. NG-27 [Bacillus sp. NG-27] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| O94163 | 2.00e-14 | 389 | 716 | 61 | 327 | Endo-1,4-beta-xylanase F1 OS=Aspergillus oryzae (strain ATCC 42149 / RIB 40) OX=510516 GN=xynF1 PE=1 SV=1 |
| O59859 | 2.80e-13 | 401 | 716 | 77 | 327 | Endo-1,4-beta-xylanase OS=Aspergillus aculeatus OX=5053 GN=xynIA PE=3 SV=1 |
| Q96VB6 | 4.83e-13 | 453 | 716 | 123 | 323 | Endo-1,4-beta-xylanase F3 OS=Aspergillus oryzae (strain ATCC 42149 / RIB 40) OX=510516 GN=xynF3 PE=1 SV=1 |
| P14768 | 1.01e-12 | 435 | 719 | 344 | 611 | Endo-1,4-beta-xylanase A OS=Cellvibrio japonicus (strain Ueda107) OX=498211 GN=xynA PE=1 SV=2 |
| P07528 | 1.15e-11 | 450 | 709 | 162 | 391 | Endo-1,4-beta-xylanase A OS=Alkalihalobacillus halodurans (strain ATCC BAA-125 / DSM 18197 / FERM 7344 / JCM 9153 / C-125) OX=272558 GN=xynA PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as LIPO
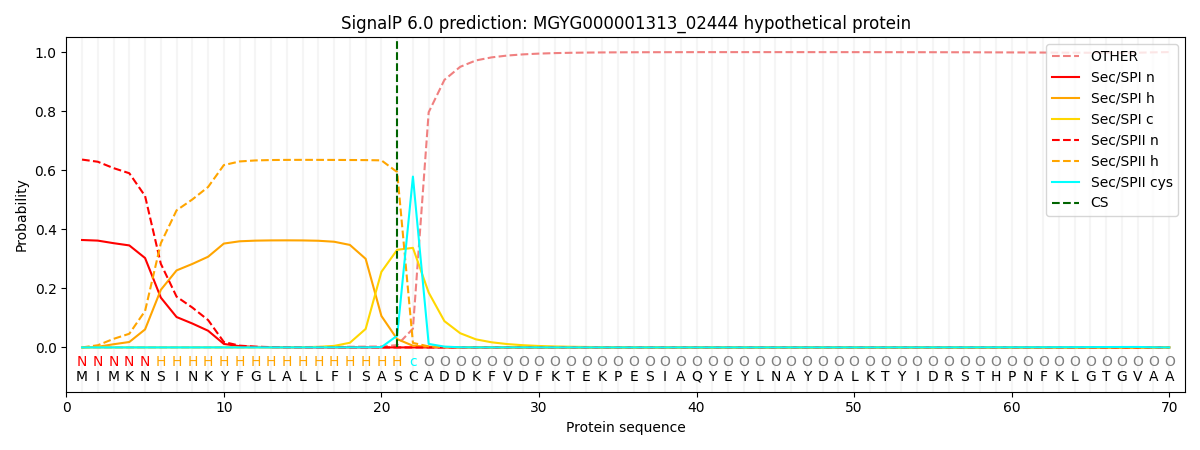
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000668 | 0.355840 | 0.642945 | 0.000175 | 0.000186 | 0.000168 |
