You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001314_01462
You are here: Home > Sequence: MGYG000001314_01462
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
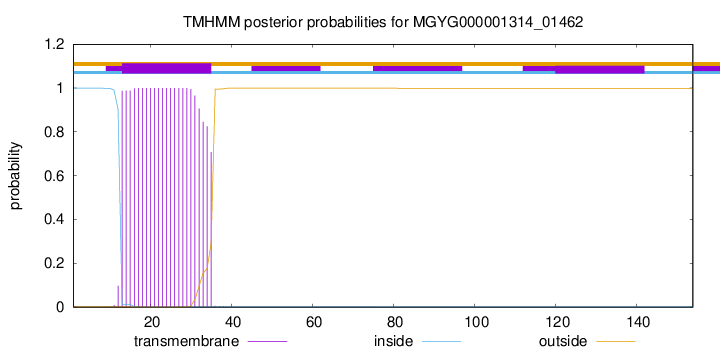
TMHMM annotations
Basic Information help
| Species | Bacteroides_F pectinophilus | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Lachnospirales; Lachnospiraceae; Bacteroides_F; Bacteroides_F pectinophilus | |||||||||||
| CAZyme ID | MGYG000001314_01462 | |||||||||||
| CAZy Family | GT51 | |||||||||||
| CAZyme Description | Penicillin-binding protein 1A | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 970742; End: 971206 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GT51 | 71 | 152 | 1.6e-26 | 0.4519774011299435 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam00912 | Transgly | 3.42e-30 | 83 | 152 | 15 | 84 | Transglycosylase. The penicillin-binding proteins are bifunctional proteins consisting of transglycosylase and transpeptidase in the N- and C-terminus respectively. The transglycosylase domain catalyzes the polymerization of murein glycan chains. |
| COG5009 | MrcA | 7.10e-27 | 60 | 152 | 47 | 138 | Membrane carboxypeptidase/penicillin-binding protein [Cell wall/membrane/envelope biogenesis]. |
| COG0744 | MrcB | 1.12e-24 | 44 | 152 | 38 | 148 | Membrane carboxypeptidase (penicillin-binding protein) [Cell wall/membrane/envelope biogenesis]. |
| PRK11636 | mrcA | 2.23e-17 | 84 | 152 | 69 | 137 | penicillin-binding protein 1a; Provisional |
| PRK00056 | mtgA | 1.44e-16 | 84 | 146 | 63 | 125 | monofunctional biosynthetic peptidoglycan transglycosylase; Provisional |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| ACR72912.1 | 1.98e-35 | 31 | 151 | 36 | 156 |
| CUH93821.1 | 1.67e-29 | 38 | 151 | 49 | 163 |
| BCN28959.1 | 2.50e-29 | 15 | 149 | 33 | 168 |
| AQP39446.1 | 2.70e-25 | 36 | 151 | 50 | 166 |
| CBL38788.1 | 5.01e-25 | 36 | 151 | 50 | 166 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3NB6_A | 1.12e-17 | 66 | 152 | 6 | 91 | Crystalstructure of Aquifex aeolicus peptidoglycan glycosyltransferase in complex with Methylphosphoryl Neryl Moenomycin [Aquifex aeolicus] |
| 2OQO_A | 1.57e-17 | 66 | 152 | 6 | 91 | Crystalstructure of a peptidoglycan glycosyltransferase from a class A PBP: insight into bacterial cell wall synthesis [Aquifex aeolicus VF5],3D3H_A Crystal structure of a complex of the peptidoglycan glycosyltransferase domain from Aquifex aeolicus and neryl moenomycin A [Aquifex aeolicus],3NB7_A Crystal structure of Aquifex Aeolicus Peptidoglycan Glycosyltransferase in complex with Decarboxylated Neryl Moenomycin [Aquifex aeolicus] |
| 5U2G_A | 5.74e-14 | 84 | 152 | 43 | 111 | 2.6Angstrom Resolution Crystal Structure of Penicillin-Binding Protein 1A from Haemophilus influenzae [Haemophilus influenzae Rd KW20],5U2G_B 2.6 Angstrom Resolution Crystal Structure of Penicillin-Binding Protein 1A from Haemophilus influenzae [Haemophilus influenzae Rd KW20] |
| 5FGZ_A | 3.60e-13 | 72 | 149 | 144 | 224 | E.coli PBP1b in complex with FPI-1465 [Escherichia coli K-12],5HL9_A E. coli PBP1b in complex with acyl-ampicillin and moenomycin [Escherichia coli K-12],5HLA_A E. coli PBP1b in complex with acyl-cephalexin and moenomycin [Escherichia coli K-12],5HLB_A E. coli PBP1b in complex with acyl-aztreonam and moenomycin [Escherichia coli K-12],5HLD_A E. coli PBP1b in complex with acyl-CENTA and moenomycin [Escherichia coli K-12],6YN0_A Structure of E. coli PBP1b with a FtsN peptide activating transglycosylase activity [Escherichia coli K-12],7LQ6_A Chain A, Penicillin-binding protein 1B [Escherichia coli K-12] |
| 3VMA_A | 3.61e-13 | 72 | 149 | 165 | 245 | CrystalStructure of the Full-Length Transglycosylase PBP1b from Escherichia coli [Escherichia coli K-12] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P40750 | 6.08e-18 | 55 | 149 | 49 | 144 | Penicillin-binding protein 4 OS=Bacillus subtilis (strain 168) OX=224308 GN=pbpD PE=1 SV=2 |
| O66874 | 1.58e-17 | 60 | 152 | 43 | 134 | Penicillin-binding protein 1A OS=Aquifex aeolicus (strain VF5) OX=224324 GN=mrcA PE=1 SV=1 |
| Q07259 | 4.14e-15 | 83 | 151 | 49 | 116 | Putative transglycosylase H16_A0665 OS=Cupriavidus necator (strain ATCC 17699 / DSM 428 / KCTC 22496 / NCIMB 10442 / H16 / Stanier 337) OX=381666 GN=H16_A0665 PE=3 SV=2 |
| Q891X1 | 5.65e-15 | 59 | 152 | 58 | 155 | Penicillin-binding protein 1A OS=Clostridium tetani (strain Massachusetts / E88) OX=212717 GN=pbpA PE=3 SV=1 |
| P39793 | 7.83e-15 | 57 | 152 | 72 | 166 | Penicillin-binding protein 1A/1B OS=Bacillus subtilis (strain 168) OX=224308 GN=ponA PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
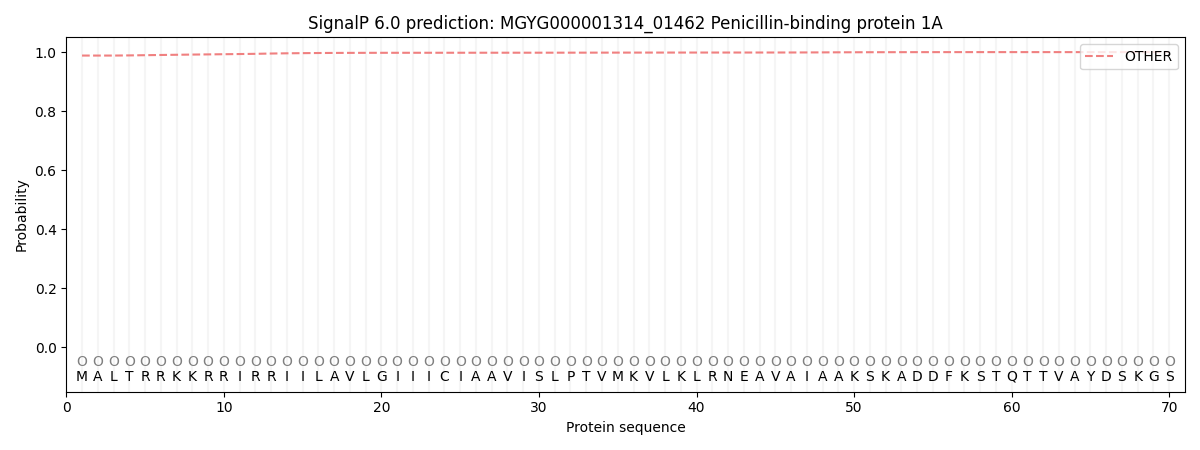
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.988411 | 0.002254 | 0.000569 | 0.000024 | 0.000011 | 0.008752 |