You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001318_00338
You are here: Home > Sequence: MGYG000001318_00338
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Enterobacter cancerogenus | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Proteobacteria; Gammaproteobacteria; Enterobacterales; Enterobacteriaceae; Enterobacter; Enterobacter cancerogenus | |||||||||||
| CAZyme ID | MGYG000001318_00338 | |||||||||||
| CAZy Family | GH20 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 135241; End: 137625 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH20 | 154 | 585 | 1.3e-119 | 0.9762611275964391 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd06570 | GH20_chitobiase-like_1 | 3.76e-175 | 159 | 597 | 1 | 311 | A functionally uncharacterized subgroup of the Glycosyl hydrolase family 20 (GH20) catalytic domain found in proteins similar to the chitobiase of Serratia marcescens, a beta-N-1,4-acetylhexosaminidase that hydrolyzes the beta-1,4-glycosidic linkages in oligomers derived from chitin. Chitin is degraded by a two step process: i) a chitinase hydrolyzes the chitin to oligosaccharides and disaccharides such as di-N-acetyl-D-glucosamine and chitobiose, ii) chitobiase then further degrades these oligomers into monomers. This subgroup lacks the C-terminal PKD (polycystic kidney disease I)-like domain found in the chitobiases. The GH20 hexosaminidases are thought to act via a catalytic mechanism in which the catalytic nucleophile is not provided by solvent or the enzyme, but by the substrate itself. |
| cd06563 | GH20_chitobiase-like | 1.01e-129 | 159 | 597 | 1 | 357 | The chitobiase of Serratia marcescens is a beta-N-1,4-acetylhexosaminidase with a glycosyl hydrolase family 20 (GH20) domain that hydrolyzes the beta-1,4-glycosidic linkages in oligomers derived from chitin. Chitin is degraded by a two step process: i) a chitinase hydrolyzes the chitin to oligosaccharides and disaccharides such as di-N-acetyl-D-glucosamine and chitobiose, ii) chitobiase then further degrades these oligomers into monomers. This GH20 domain family includes an N-acetylglucosamidase (GlcNAcase A) from Pseudoalteromonas piscicida and an N-acetylhexosaminidase (SpHex) from Streptomyces plicatus. SpHex lacks the C-terminal PKD (polycystic kidney disease I)-like domain found in the chitobiases. The GH20 hexosaminidases are thought to act via a catalytic mechanism in which the catalytic nucleophile is not provided by solvent or the enzyme, but by the substrate itself. |
| pfam00728 | Glyco_hydro_20 | 4.35e-120 | 159 | 584 | 1 | 345 | Glycosyl hydrolase family 20, catalytic domain. This domain has a TIM barrel fold. |
| cd06562 | GH20_HexA_HexB-like | 2.07e-108 | 159 | 600 | 1 | 340 | Beta-N-acetylhexosaminidases catalyze the removal of beta-1,4-linked N-acetyl-D-hexosamine residues from the non-reducing ends of N-acetyl-beta-D-hexosaminides including N-acetylglucosides and N-acetylgalactosides. The hexA and hexB genes encode the alpha- and beta-subunits of the two major beta-N-acetylhexosaminidase isoenzymes, N-acetyl-beta-D-hexosaminidase A (HexA) and beta-N-acetylhexosaminidase B (HexB). Both the alpha and the beta catalytic subunits have a TIM-barrel fold and belong to the glycosyl hydrolase family 20 (GH20). The HexA enzyme is a heterodimer containing one alpha and one beta subunit while the HexB enzyme is a homodimer containing two beta-subunits. Hexosaminidase mutations cause an inability to properly hydrolyze certain sphingolipids which accumulate in lysosomes within the brain, resulting in the lipid storage disorders Tay-Sachs and Sandhoff. Mutations in the alpha subunit cause in a deficiency in the HexA enzyme and result in Tay-Sachs, mutations in the beta-subunit cause in a deficiency in both HexA and HexB enzymes and result in Sandhoff disease. In both disorders GM(2) gangliosides accumulate in lysosomes. The GH20 hexosaminidases are thought to act via a catalytic mechanism in which the catalytic nucleophile is not provided by solvent or the enzyme, but by the substrate itself. |
| cd02742 | GH20_hexosaminidase | 1.03e-83 | 161 | 583 | 1 | 303 | Beta-N-acetylhexosaminidases of glycosyl hydrolase family 20 (GH20) catalyze the removal of beta-1,4-linked N-acetyl-D-hexosamine residues from the non-reducing ends of N-acetyl-beta-D-hexosaminides including N-acetylglucosides and N-acetylgalactosides. These enzymes are broadly distributed in microorganisms, plants and animals, and play roles in various key physiological and pathological processes. These processes include cell structural integrity, energy storage, cellular signaling, fertilization, pathogen defense, viral penetration, the development of carcinomas, inflammatory events and lysosomal storage disorders. The GH20 enzymes include the eukaryotic beta-N-acetylhexosaminidases A and B, the bacterial chitobiases, dispersin B, and lacto-N-biosidase. The GH20 hexosaminidases are thought to act via a catalytic mechanism in which the catalytic nucleophile is not provided by the solvent or the enzyme, but by the substrate itself. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| CAD5352166.1 | 0.0 | 1 | 794 | 1 | 794 |
| QZY37436.1 | 0.0 | 1 | 794 | 1 | 794 |
| AUJ83146.1 | 0.0 | 1 | 794 | 1 | 794 |
| QGG09778.1 | 0.0 | 1 | 794 | 1 | 794 |
| ARZ78388.1 | 0.0 | 1 | 794 | 1 | 794 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6YHH_A | 1.07e-114 | 24 | 769 | 6 | 658 | X-rayStructure of Flavobacterium johnsoniae chitobiase (FjGH20) [Flavobacterium johnsoniae UW101],6YHH_B X-ray Structure of Flavobacterium johnsoniae chitobiase (FjGH20) [Flavobacterium johnsoniae UW101] |
| 7CBN_A | 2.95e-70 | 107 | 384 | 78 | 377 | Crystalstructure of beta-N-acetylhexosaminidase Am0868 from Akkermansia muciniphila [Akkermansia muciniphila ATCC BAA-835],7CBO_A Crystal structure of beta-N-acetylhexosaminidase Am0868 from Akkermansia muciniphila in complex with GlcNAc [Akkermansia muciniphila ATCC BAA-835] |
| 6Q63_A | 1.14e-64 | 73 | 426 | 78 | 448 | BT0459[Bacteroides thetaiotaomicron],6Q63_B BT0459 [Bacteroides thetaiotaomicron],6Q63_C BT0459 [Bacteroides thetaiotaomicron] |
| 3RCN_A | 3.14e-60 | 73 | 410 | 53 | 413 | CrystalStructure of Beta-N-Acetylhexosaminidase from Arthrobacter aurescens [Paenarthrobacter aurescens TC1] |
| 1NOU_A | 7.16e-60 | 105 | 597 | 96 | 480 | Nativehuman lysosomal beta-hexosaminidase isoform B [Homo sapiens],1NOU_B Native human lysosomal beta-hexosaminidase isoform B [Homo sapiens],1NOW_A Human lysosomal beta-hexosaminidase isoform B in complex with (2R,3R,4S,5R)-2-Acetamido-3,4-Dihydroxy-5-Hydroxymethyl-Piperidinium Chloride (GalNAc-isofagomine) [Homo sapiens],1NOW_B Human lysosomal beta-hexosaminidase isoform B in complex with (2R,3R,4S,5R)-2-Acetamido-3,4-Dihydroxy-5-Hydroxymethyl-Piperidinium Chloride (GalNAc-isofagomine) [Homo sapiens],2GJX_B Crystallographic structure of human beta-Hexosaminidase A [Homo sapiens],2GJX_C Crystallographic structure of human beta-Hexosaminidase A [Homo sapiens],2GJX_F Crystallographic structure of human beta-Hexosaminidase A [Homo sapiens],2GJX_G Crystallographic structure of human beta-Hexosaminidase A [Homo sapiens] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P49008 | 3.80e-75 | 31 | 409 | 39 | 432 | Beta-hexosaminidase OS=Porphyromonas gingivalis (strain ATCC BAA-308 / W83) OX=242619 GN=nahA PE=3 SV=2 |
| B2UQG6 | 2.09e-69 | 107 | 384 | 97 | 396 | Beta-hexosaminidase Amuc_0868 OS=Akkermansia muciniphila (strain ATCC BAA-835 / DSM 22959 / JCM 33894 / BCRC 81048 / CCUG 64013 / CIP 107961 / Muc) OX=349741 GN=Amuc_0868 PE=1 SV=1 |
| P96155 | 2.55e-64 | 19 | 412 | 133 | 535 | Beta-hexosaminidase OS=Vibrio furnissii OX=29494 GN=exoI PE=1 SV=1 |
| P20060 | 4.56e-62 | 85 | 597 | 104 | 508 | Beta-hexosaminidase subunit beta OS=Mus musculus OX=10090 GN=Hexb PE=1 SV=2 |
| P13723 | 7.95e-62 | 106 | 597 | 100 | 483 | Beta-hexosaminidase subunit A1 OS=Dictyostelium discoideum OX=44689 GN=hexa1 PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
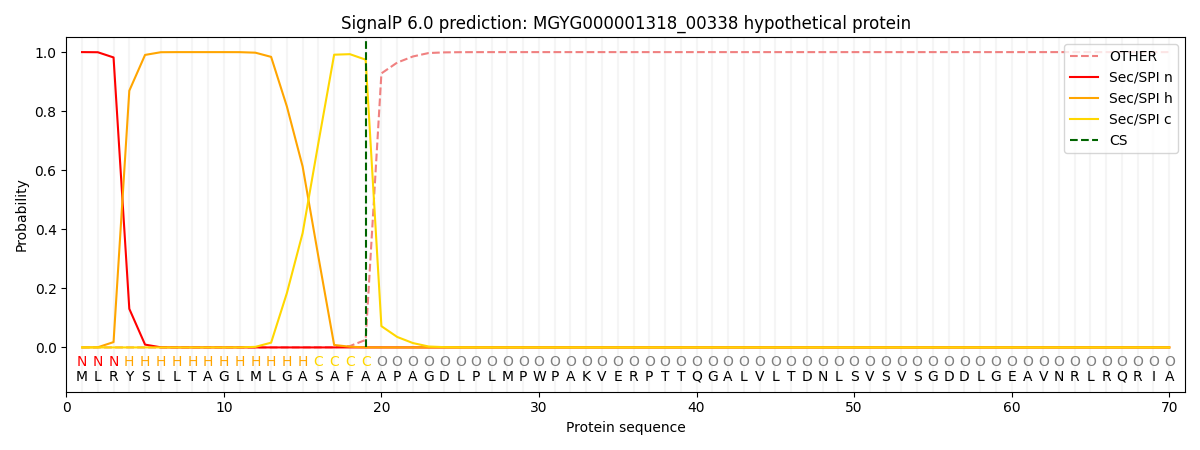
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000396 | 0.998762 | 0.000189 | 0.000238 | 0.000213 | 0.000178 |
