You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001332_01594
You are here: Home > Sequence: MGYG000001332_01594
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Massilioclostridium methylpentosum | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Oscillospirales; Ruminococcaceae; Massilioclostridium; Massilioclostridium methylpentosum | |||||||||||
| CAZyme ID | MGYG000001332_01594 | |||||||||||
| CAZy Family | GH106 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 31225; End: 34512 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH106 | 38 | 836 | 2.6e-74 | 0.8288834951456311 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam17132 | Glyco_hydro_106 | 1.28e-13 | 291 | 737 | 436 | 835 | alpha-L-rhamnosidase. |
| pfam02837 | Glyco_hydro_2_N | 9.68e-06 | 915 | 986 | 61 | 133 | Glycosyl hydrolases family 2, sugar binding domain. This family contains beta-galactosidase, beta-mannosidase and beta-glucuronidase activities and has a jelly-roll fold. The domain binds the sugar moiety during the sugar-hydrolysis reaction. |
| PRK10340 | ebgA | 0.003 | 915 | 986 | 105 | 177 | cryptic beta-D-galactosidase subunit alpha; Reviewed |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QGH70002.1 | 3.49e-239 | 32 | 1029 | 85 | 1116 |
| QYN20448.1 | 9.43e-212 | 11 | 1029 | 24 | 1121 |
| SDG73971.1 | 6.15e-208 | 41 | 1026 | 10 | 960 |
| QHO90364.1 | 1.32e-199 | 37 | 1022 | 32 | 1043 |
| QNP57182.1 | 1.52e-198 | 42 | 1020 | 12 | 971 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6Q2F_A | 7.01e-07 | 197 | 539 | 405 | 734 | Structureof Rhamnosidase from Novosphingobium sp. PP1Y [Novosphingobium sp. PP1Y] |
Swiss-Prot Hits help
SignalP and Lipop Annotations help
This protein is predicted as SP
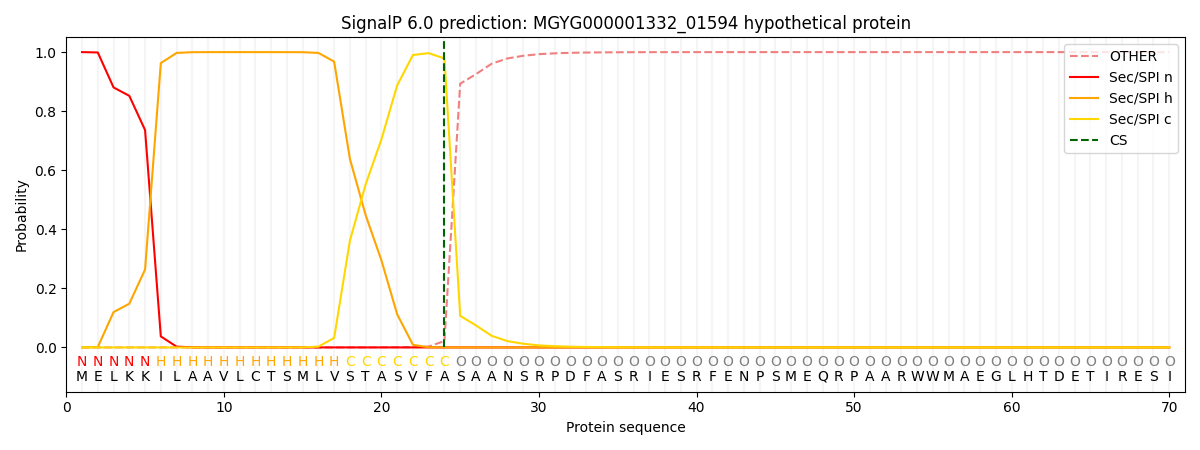
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000433 | 0.998713 | 0.000233 | 0.000217 | 0.000192 | 0.000165 |

