You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001334_00021
You are here: Home > Sequence: MGYG000001334_00021
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Ligilactobacillus ruminis | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes; Bacilli; Lactobacillales; Lactobacillaceae; Ligilactobacillus; Ligilactobacillus ruminis | |||||||||||
| CAZyme ID | MGYG000001334_00021 | |||||||||||
| CAZy Family | GH73 | |||||||||||
| CAZyme Description | Cell division suppressor protein YneA | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 3580; End: 5979 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH73 | 171 | 307 | 8.4e-30 | 0.984375 |
| CBM50 | 420 | 461 | 2.2e-16 | 0.95 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| PRK06347 | PRK06347 | 2.53e-84 | 2 | 667 | 3 | 592 | 1,4-beta-N-acetylmuramoylhydrolase. |
| COG1705 | FlgJ | 1.10e-46 | 160 | 313 | 40 | 190 | Flagellum-specific peptidoglycan hydrolase FlgJ [Cell wall/membrane/envelope biogenesis, Cell motility]. |
| PRK06347 | PRK06347 | 8.63e-40 | 583 | 796 | 291 | 521 | 1,4-beta-N-acetylmuramoylhydrolase. |
| PRK08581 | PRK08581 | 1.94e-27 | 160 | 311 | 317 | 472 | amidase domain-containing protein. |
| PRK05684 | flgJ | 3.44e-27 | 128 | 304 | 118 | 296 | flagellar assembly peptidoglycan hydrolase FlgJ. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| AEN78697.1 | 0.0 | 1 | 799 | 1 | 725 |
| ASR40346.1 | 2.45e-201 | 1 | 796 | 1 | 773 |
| QHQ69680.1 | 1.58e-199 | 1 | 798 | 1 | 760 |
| AWZ38490.1 | 8.14e-191 | 1 | 798 | 1 | 760 |
| AWZ40523.1 | 8.14e-191 | 1 | 798 | 1 | 760 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3FI7_A | 2.61e-20 | 164 | 311 | 31 | 183 | CrystalStructure of the autolysin Auto (Lmo1076) from Listeria monocytogenes, catalytic domain [Listeria monocytogenes EGD-e] |
| 5T1Q_A | 5.31e-18 | 154 | 311 | 51 | 212 | ChainA, N-acetylmuramoyl-L-alanine amidase domain-containing protein SAOUHSC_02979 [Staphylococcus aureus subsp. aureus NCTC 8325],5T1Q_B Chain B, N-acetylmuramoyl-L-alanine amidase domain-containing protein SAOUHSC_02979 [Staphylococcus aureus subsp. aureus NCTC 8325],5T1Q_C Chain C, N-acetylmuramoyl-L-alanine amidase domain-containing protein SAOUHSC_02979 [Staphylococcus aureus subsp. aureus NCTC 8325],5T1Q_D Chain D, N-acetylmuramoyl-L-alanine amidase domain-containing protein SAOUHSC_02979 [Staphylococcus aureus subsp. aureus NCTC 8325] |
| 3VWO_A | 1.57e-16 | 164 | 309 | 2 | 150 | Crystalstructure of peptidoglycan hydrolase mutant from Sphingomonas sp. A1 [Sphingomonas sp. A1] |
| 2ZYC_A | 1.96e-16 | 164 | 309 | 3 | 151 | ChainA, Peptidoglycan hydrolase FlgJ [Sphingomonas sp. A1] |
| 5DN5_A | 7.16e-16 | 164 | 304 | 4 | 147 | Structureof a C-terminally truncated glycoside hydrolase domain from Salmonella typhimurium FlgJ [Salmonella enterica subsp. enterica serovar Typhimurium str. LT2],5DN5_B Structure of a C-terminally truncated glycoside hydrolase domain from Salmonella typhimurium FlgJ [Salmonella enterica subsp. enterica serovar Typhimurium str. LT2],5DN5_C Structure of a C-terminally truncated glycoside hydrolase domain from Salmonella typhimurium FlgJ [Salmonella enterica subsp. enterica serovar Typhimurium str. LT2] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P37710 | 1.60e-91 | 147 | 732 | 160 | 736 | Autolysin OS=Enterococcus faecalis (strain ATCC 700802 / V583) OX=226185 GN=EF_0799 PE=1 SV=2 |
| P39046 | 8.75e-90 | 148 | 732 | 40 | 664 | Muramidase-2 OS=Enterococcus hirae (strain ATCC 9790 / DSM 20160 / JCM 8729 / LMG 6399 / NBRC 3181 / NCIMB 6459 / NCDO 1258 / NCTC 12367 / WDCM 00089 / R) OX=768486 GN=EHR_05900 PE=1 SV=1 |
| A2RHZ5 | 2.42e-51 | 153 | 525 | 55 | 434 | Probable N-acetylmuramidase OS=Lactococcus lactis subsp. cremoris (strain MG1363) OX=416870 GN=acmA PE=3 SV=1 |
| P0C2T5 | 3.03e-50 | 153 | 525 | 55 | 434 | Probable N-acetylmuramidase OS=Lactococcus lactis subsp. cremoris OX=1359 GN=acmA PE=3 SV=1 |
| Q9CIT4 | 5.96e-50 | 153 | 525 | 55 | 436 | Probable N-acetylmuramidase OS=Lactococcus lactis subsp. lactis (strain IL1403) OX=272623 GN=acmA PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
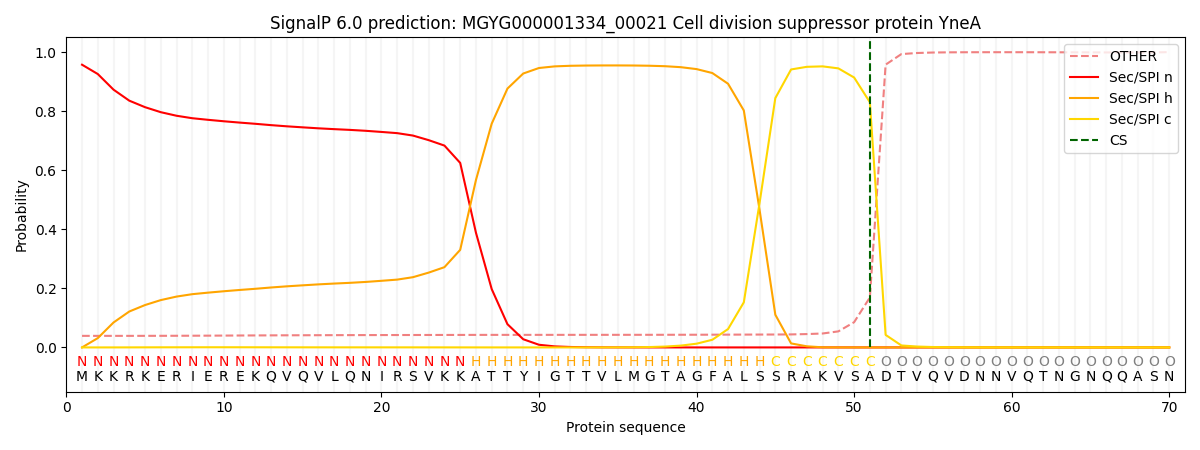
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.045555 | 0.948509 | 0.003972 | 0.001278 | 0.000371 | 0.000305 |
