You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001336_01020
You are here: Home > Sequence: MGYG000001336_01020
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
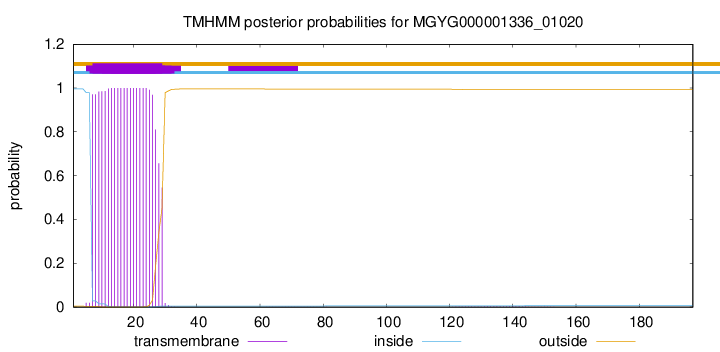
TMHMM annotations
Basic Information help
| Species | Limosilactobacillus reuteri_E | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes; Bacilli; Lactobacillales; Lactobacillaceae; Limosilactobacillus; Limosilactobacillus reuteri_E | |||||||||||
| CAZyme ID | MGYG000001336_01020 | |||||||||||
| CAZy Family | GH73 | |||||||||||
| CAZyme Description | Exo-glucosaminidase LytG | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 982717; End: 983310 Strand: - | |||||||||||
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG1705 | FlgJ | 2.24e-55 | 1 | 197 | 1 | 191 | Flagellum-specific peptidoglycan hydrolase FlgJ [Cell wall/membrane/envelope biogenesis, Cell motility]. |
| PRK05684 | flgJ | 3.68e-31 | 46 | 188 | 154 | 297 | flagellar assembly peptidoglycan hydrolase FlgJ. |
| NF038016 | sporang_Gsm | 4.27e-30 | 47 | 194 | 163 | 312 | sporangiospore maturation cell wall hydrolase GsmA. The peptidoglycan-hydrolyzing enzyme GsmA occurs in some sporangia-forming members of the Actinobacteria, such as Actinoplanes missouriensis, and is required for proper separation of spores. GsmA proteins have one or two SH3 domains N-terminal to the hydrolase domain. |
| smart00047 | LYZ2 | 6.56e-30 | 46 | 194 | 10 | 147 | Lysozyme subfamily 2. Eubacterial enzymes distantly related to eukaryotic lysozymes. |
| TIGR02541 | flagell_FlgJ | 7.76e-29 | 46 | 187 | 150 | 292 | flagellar rod assembly protein/muramidase FlgJ. The N-terminal region of this protein acts directly in flagellar rod assembly. The C-terminal region is a flagellum-specific muramidase (peptidoglycan hydrolase) required for formation of the outer membrane L ring. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QLL76945.1 | 3.76e-140 | 1 | 197 | 1 | 197 |
| AAY86786.1 | 3.76e-140 | 1 | 197 | 1 | 197 |
| QLQ62841.1 | 3.76e-140 | 1 | 197 | 1 | 197 |
| AEI57215.1 | 3.76e-140 | 1 | 197 | 1 | 197 |
| ANU52081.1 | 8.51e-137 | 1 | 197 | 1 | 197 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3FI7_A | 1.01e-21 | 47 | 194 | 33 | 183 | CrystalStructure of the autolysin Auto (Lmo1076) from Listeria monocytogenes, catalytic domain [Listeria monocytogenes EGD-e] |
| 5DN5_A | 8.38e-17 | 46 | 187 | 5 | 147 | Structureof a C-terminally truncated glycoside hydrolase domain from Salmonella typhimurium FlgJ [Salmonella enterica subsp. enterica serovar Typhimurium str. LT2],5DN5_B Structure of a C-terminally truncated glycoside hydrolase domain from Salmonella typhimurium FlgJ [Salmonella enterica subsp. enterica serovar Typhimurium str. LT2],5DN5_C Structure of a C-terminally truncated glycoside hydrolase domain from Salmonella typhimurium FlgJ [Salmonella enterica subsp. enterica serovar Typhimurium str. LT2] |
| 5DN4_A | 1.16e-16 | 46 | 187 | 5 | 147 | Structureof the glycoside hydrolase domain from Salmonella typhimurium FlgJ [Salmonella enterica subsp. enterica serovar Typhimurium str. LT2] |
| 5T1Q_A | 1.78e-16 | 46 | 195 | 62 | 213 | ChainA, N-acetylmuramoyl-L-alanine amidase domain-containing protein SAOUHSC_02979 [Staphylococcus aureus subsp. aureus NCTC 8325],5T1Q_B Chain B, N-acetylmuramoyl-L-alanine amidase domain-containing protein SAOUHSC_02979 [Staphylococcus aureus subsp. aureus NCTC 8325],5T1Q_C Chain C, N-acetylmuramoyl-L-alanine amidase domain-containing protein SAOUHSC_02979 [Staphylococcus aureus subsp. aureus NCTC 8325],5T1Q_D Chain D, N-acetylmuramoyl-L-alanine amidase domain-containing protein SAOUHSC_02979 [Staphylococcus aureus subsp. aureus NCTC 8325] |
| 3VWO_A | 1.90e-13 | 47 | 192 | 4 | 150 | Crystalstructure of peptidoglycan hydrolase mutant from Sphingomonas sp. A1 [Sphingomonas sp. A1] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| O32083 | 9.25e-34 | 1 | 196 | 6 | 200 | Exo-glucosaminidase LytG OS=Bacillus subtilis (strain 168) OX=224308 GN=lytG PE=1 SV=1 |
| P39046 | 5.54e-17 | 29 | 194 | 43 | 214 | Muramidase-2 OS=Enterococcus hirae (strain ATCC 9790 / DSM 20160 / JCM 8729 / LMG 6399 / NBRC 3181 / NCIMB 6459 / NCDO 1258 / NCTC 12367 / WDCM 00089 / R) OX=768486 GN=EHR_05900 PE=1 SV=1 |
| P58231 | 9.51e-16 | 47 | 187 | 152 | 293 | Peptidoglycan hydrolase FlgJ OS=Escherichia coli O157:H7 OX=83334 GN=flgJ PE=3 SV=1 |
| P75942 | 9.51e-16 | 47 | 187 | 152 | 293 | Peptidoglycan hydrolase FlgJ OS=Escherichia coli (strain K12) OX=83333 GN=flgJ PE=3 SV=1 |
| Q9CIT4 | 9.78e-16 | 30 | 194 | 52 | 214 | Probable N-acetylmuramidase OS=Lactococcus lactis subsp. lactis (strain IL1403) OX=272623 GN=acmA PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
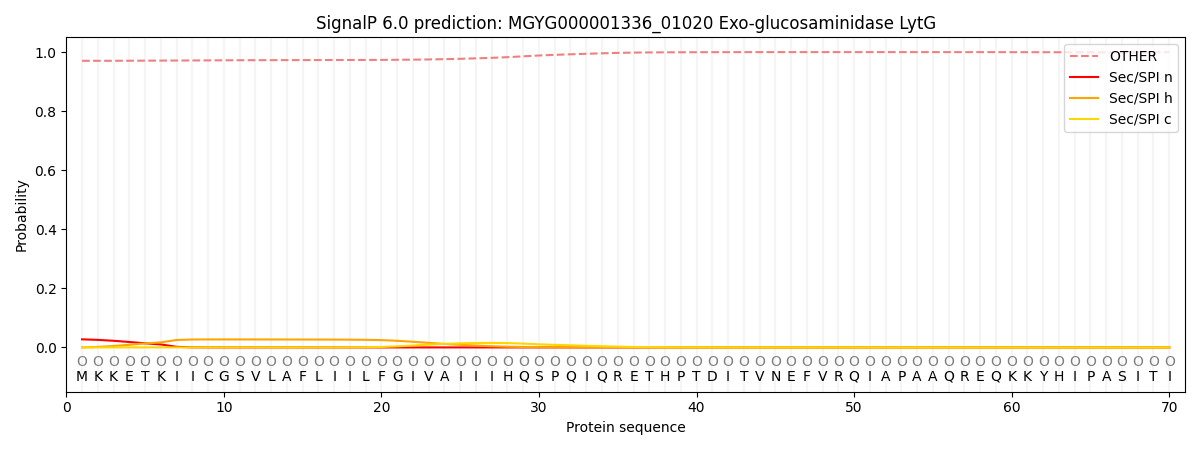
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.971934 | 0.025286 | 0.000484 | 0.000141 | 0.000094 | 0.002073 |