You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001338_00199
You are here: Home > Sequence: MGYG000001338_00199
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Blautia_A wexlerae_A | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Lachnospirales; Lachnospiraceae; Blautia_A; Blautia_A wexlerae_A | |||||||||||
| CAZyme ID | MGYG000001338_00199 | |||||||||||
| CAZy Family | GH3 | |||||||||||
| CAZyme Description | Beta-hexosaminidase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 201031; End: 203151 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH3 | 37 | 277 | 6.3e-70 | 0.9814814814814815 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| PLN03080 | PLN03080 | 1.16e-141 | 14 | 677 | 52 | 764 | Probable beta-xylosidase; Provisional |
| PRK15098 | PRK15098 | 6.39e-128 | 40 | 695 | 101 | 763 | beta-glucosidase BglX. |
| pfam01915 | Glyco_hydro_3_C | 8.69e-72 | 346 | 581 | 2 | 216 | Glycosyl hydrolase family 3 C-terminal domain. This domain is involved in catalysis and may be involved in binding beta-glucan. This domain is found associated with pfam00933. |
| COG1472 | BglX | 1.13e-64 | 40 | 413 | 57 | 397 | Periplasmic beta-glucosidase and related glycosidases [Carbohydrate transport and metabolism]. |
| pfam00933 | Glyco_hydro_3 | 2.59e-47 | 26 | 307 | 46 | 315 | Glycosyl hydrolase family 3 N terminal domain. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QIB55396.1 | 0.0 | 1 | 706 | 1 | 706 |
| QMW76741.1 | 0.0 | 1 | 706 | 1 | 706 |
| QBE95420.1 | 0.0 | 1 | 706 | 1 | 706 |
| QJU15895.1 | 0.0 | 1 | 704 | 1 | 704 |
| ASU31110.1 | 0.0 | 1 | 704 | 1 | 704 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 7VC7_A | 5.16e-102 | 14 | 696 | 25 | 731 | ChainA, xylan 1,4-beta-xylosidase [Phanerodontia chrysosporium] |
| 7VC6_A | 5.16e-102 | 14 | 696 | 25 | 731 | ChainA, xylan 1,4-beta-xylosidase [Phanerodontia chrysosporium] |
| 5Z87_A | 3.72e-101 | 40 | 695 | 120 | 780 | ChainA, EmGH1 [Aurantiacibacter marinus],5Z87_B Chain B, EmGH1 [Aurantiacibacter marinus] |
| 6Q7I_A | 1.76e-94 | 14 | 698 | 52 | 755 | GH3exo-beta-xylosidase (XlnD) [Aspergillus nidulans FGSC A4],6Q7I_B GH3 exo-beta-xylosidase (XlnD) [Aspergillus nidulans FGSC A4],6Q7J_A Chain A, Exo-1,4-beta-xylosidase xlnD [Aspergillus nidulans FGSC A4],6Q7J_B Chain B, Exo-1,4-beta-xylosidase xlnD [Aspergillus nidulans FGSC A4] |
| 5A7M_A | 6.86e-90 | 14 | 698 | 52 | 757 | ChainA, BETA-XYLOSIDASE [Trichoderma reesei],5A7M_B Chain B, BETA-XYLOSIDASE [Trichoderma reesei] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| D5EY15 | 5.31e-147 | 11 | 695 | 31 | 858 | Xylan 1,4-beta-xylosidase OS=Prevotella ruminicola (strain ATCC 19189 / JCM 8958 / 23) OX=264731 GN=xyl3A PE=1 SV=1 |
| Q9SGZ5 | 1.59e-124 | 13 | 582 | 47 | 626 | Probable beta-D-xylosidase 7 OS=Arabidopsis thaliana OX=3702 GN=BXL7 PE=2 SV=2 |
| Q9FLG1 | 1.35e-123 | 18 | 677 | 73 | 769 | Beta-D-xylosidase 4 OS=Arabidopsis thaliana OX=3702 GN=BXL4 PE=1 SV=1 |
| Q9LXD6 | 6.47e-121 | 18 | 683 | 63 | 763 | Beta-D-xylosidase 3 OS=Arabidopsis thaliana OX=3702 GN=BXL3 PE=1 SV=1 |
| Q9LJN4 | 1.11e-120 | 14 | 683 | 51 | 758 | Probable beta-D-xylosidase 5 OS=Arabidopsis thaliana OX=3702 GN=BXL5 PE=2 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
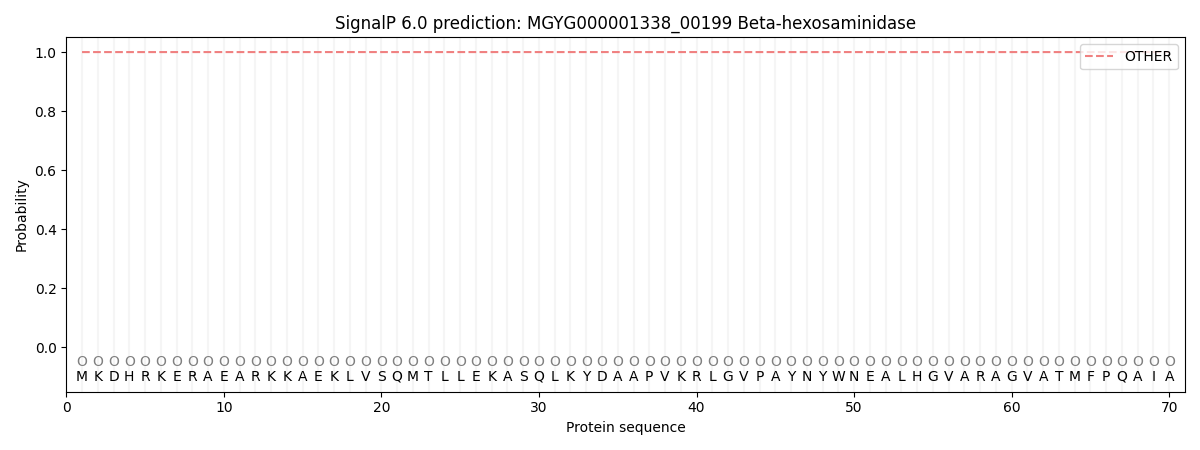
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000064 | 0.000006 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
