You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001343_00619
You are here: Home > Sequence: MGYG000001343_00619
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Acinetobacter junii | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Proteobacteria; Gammaproteobacteria; Pseudomonadales; Moraxellaceae; Acinetobacter; Acinetobacter junii | |||||||||||
| CAZyme ID | MGYG000001343_00619 | |||||||||||
| CAZy Family | GH23 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 2868; End: 3503 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH23 | 97 | 205 | 1.3e-24 | 0.8296296296296296 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd00254 | LT-like | 1.03e-20 | 103 | 185 | 1 | 87 | lytic transglycosylase(LT)-like domain. Members include the soluble and insoluble membrane-bound LTs in bacteria and LTs in bacteriophage lambda. LTs catalyze the cleavage of the beta-1,4-glycosidic bond between N-acetylmuramic acid (MurNAc) and N-acetyl-D-glucosamine (GlcNAc), as do "goose-type" lysozymes. However, in addition to this, they also make a new glycosidic bond with the C6 hydroxyl group of the same muramic acid residue. |
| cd16896 | LT_Slt70-like | 2.65e-20 | 92 | 181 | 8 | 107 | uncharacterized lytic transglycosylase subfamily with similarity to Slt70. Uncharacterized lytic transglycosylase (LT) with a conserved sequence pattern suggesting similarity to the Slt70, a 70kda soluble lytic transglycosylase which also has an N-terminal U-shaped U-domain and a linker L-domain. LTs catalyze the cleavage of the beta-1,4-glycosidic bond between N-acetylmuramic acid (MurNAc) and N-acetyl-D-glucosamine (GlcNAc), as do "goose-type" lysozymes. However, in addition to this, they also make a new glycosidic bond with the C6 hydroxyl group of the same muramic acid residue. |
| cd13401 | Slt70-like | 4.21e-19 | 89 | 181 | 7 | 109 | 70kDa soluble lytic transglycosylase (Slt70) and similar proteins. Catalytic domain of the 70kda soluble lytic transglycosylase (LT)-like proteins, which also have an N-terminal U-shaped U-domain and a linker L-domain. LTs catalyze the cleavage of the beta-1,4-glycosidic bond between N-acetylmuramic acid (MurNAc) and N-acetyl-D-glucosamine (GlcNAc), as do "goose-type" lysozymes. However, in addition to this, they also make a new glycosidic bond with the C6 hydroxyl group of the same muramic acid residue. Proteins similar to this family include the soluble and insoluble membrane-bound LTs in bacteria and the LTs in bacteriophage lambda. |
| pfam01464 | SLT | 8.18e-19 | 96 | 180 | 5 | 96 | Transglycosylase SLT domain. This family is distantly related to pfam00062. Members are found in phages, type II, type III and type IV secretion systems. |
| cd13400 | LT_IagB-like | 8.51e-14 | 99 | 203 | 1 | 106 | Escherichia coli invasion protein IagB and similar proteins. Lytic transglycosylase-like protein, similar to Escherichia coli invasion protein IagB. IagB is encoded within a pathogenicity island in Salmonella enterica and has been shown to degrade polymeric peptidoglycan. IagB-like invasion proteins are implicated in the invasion of eukaryotic host cells by bacteria. Lytic transglycosylase (LT) catalyzes the cleavage of the beta-1,4-glycosidic bond between N-acetylmuramic acid (MurNAc) and N-acetyl-D-glucosamine (GlcNAc), as do "goose-type" lysozymes. However, in addition to this, they also make a new glycosidic bond with the C6 hydroxyl group of the same muramic acid residue. Members of this family resemble the soluble and insoluble membrane-bound LTs in bacteria and the LTs in bacteriophage lambda. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QQV67578.1 | 4.16e-144 | 1 | 211 | 1 | 211 |
| ATU46918.1 | 2.74e-115 | 40 | 211 | 1 | 172 |
| QXR12240.1 | 2.74e-115 | 40 | 211 | 1 | 172 |
| QUS51557.1 | 2.74e-115 | 40 | 211 | 1 | 172 |
| QXR29201.1 | 2.25e-114 | 40 | 211 | 1 | 172 |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| O31976 | 1.35e-11 | 85 | 180 | 1419 | 1515 | SPbeta prophage-derived uncharacterized transglycosylase YomI OS=Bacillus subtilis (strain 168) OX=224308 GN=yomI PE=3 SV=2 |
| O64046 | 1.35e-11 | 85 | 180 | 1419 | 1515 | Probable tape measure protein OS=Bacillus phage SPbeta OX=66797 GN=yomI PE=3 SV=1 |
| O31608 | 6.82e-09 | 92 | 185 | 64 | 158 | Putative murein lytic transglycosylase YjbJ OS=Bacillus subtilis (strain 168) OX=224308 GN=yjbJ PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
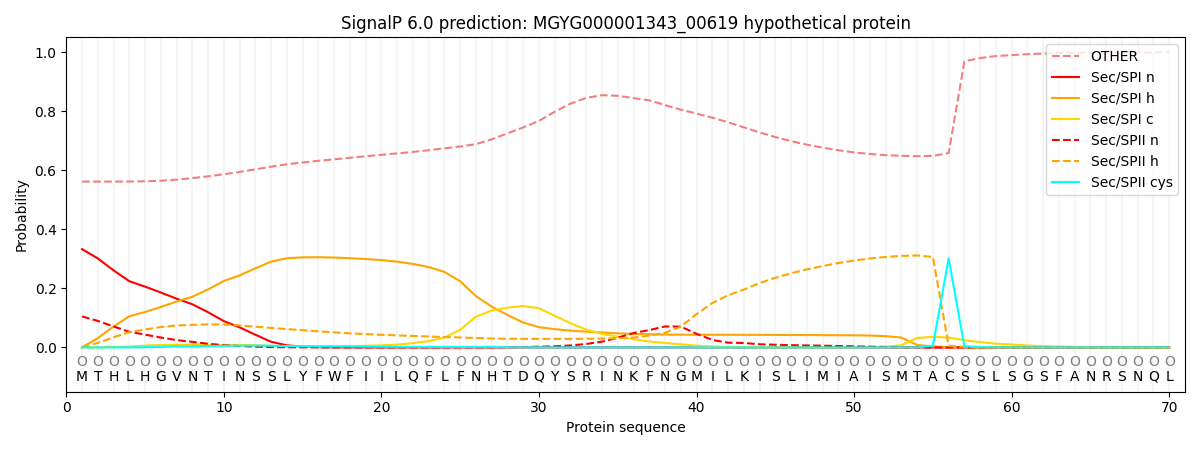
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.562836 | 0.210092 | 0.226057 | 0.000116 | 0.000185 | 0.000699 |

