You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001345_00575
You are here: Home > Sequence: MGYG000001345_00575
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Bacteroides xylanisolvens | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Bacteroidaceae; Bacteroides; Bacteroides xylanisolvens | |||||||||||
| CAZyme ID | MGYG000001345_00575 | |||||||||||
| CAZy Family | GH3 | |||||||||||
| CAZyme Description | Beta-hexosaminidase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 96771; End: 99782 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH3 | 109 | 333 | 3.4e-57 | 0.9814814814814815 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG1472 | BglX | 4.16e-77 | 57 | 430 | 4 | 360 | Periplasmic beta-glucosidase and related glycosidases [Carbohydrate transport and metabolism]. |
| pfam00933 | Glyco_hydro_3 | 2.04e-75 | 58 | 368 | 4 | 315 | Glycosyl hydrolase family 3 N terminal domain. |
| PRK03642 | PRK03642 | 4.86e-59 | 581 | 1001 | 30 | 432 | putative periplasmic esterase; Provisional |
| COG1680 | AmpC | 3.48e-45 | 592 | 977 | 36 | 367 | CubicO group peptidase, beta-lactamase class C family [Defense mechanisms]. |
| pfam00144 | Beta-lactamase | 2.50e-43 | 594 | 986 | 1 | 321 | Beta-lactamase. This family appears to be distantly related to pfam00905 and PF00768 D-alanyl-D-alanine carboxypeptidase. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QUR46065.1 | 0.0 | 1 | 1003 | 1 | 1003 |
| CBK66838.1 | 0.0 | 1 | 1003 | 1 | 1003 |
| QRN00095.1 | 0.0 | 1 | 1003 | 1 | 1003 |
| QUT31090.1 | 0.0 | 1 | 1003 | 1 | 1003 |
| QDH53003.1 | 0.0 | 1 | 1003 | 1 | 1003 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6K5J_A | 6.80e-62 | 54 | 413 | 11 | 379 | Structureof a glycoside hydrolase family 3 beta-N-acetylglucosaminidase from Paenibacillus sp. str. FPU-7 [Paenibacillaceae] |
| 3BMX_A | 3.70e-58 | 48 | 466 | 36 | 503 | Beta-N-hexosaminidase(YbbD) from Bacillus subtilis [Bacillus subtilis],3BMX_B Beta-N-hexosaminidase (YbbD) from Bacillus subtilis [Bacillus subtilis],3NVD_A Structure of YBBD in complex with pugnac [Bacillus subtilis],3NVD_B Structure of YBBD in complex with pugnac [Bacillus subtilis] |
| 3LK6_A | 1.08e-57 | 48 | 466 | 10 | 477 | ChainA, Lipoprotein ybbD [Bacillus subtilis],3LK6_B Chain B, Lipoprotein ybbD [Bacillus subtilis],3LK6_C Chain C, Lipoprotein ybbD [Bacillus subtilis],3LK6_D Chain D, Lipoprotein ybbD [Bacillus subtilis] |
| 4GYJ_A | 1.89e-57 | 48 | 466 | 40 | 507 | Crystalstructure of mutant (D318N) bacillus subtilis family 3 glycoside hydrolase (nagz) in complex with glcnac-murnac (space group P1) [Bacillus subtilis subsp. subtilis str. 168],4GYJ_B Crystal structure of mutant (D318N) bacillus subtilis family 3 glycoside hydrolase (nagz) in complex with glcnac-murnac (space group P1) [Bacillus subtilis subsp. subtilis str. 168],4GYK_A Crystal structure of mutant (D318N) bacillus subtilis family 3 glycoside hydrolase (nagz) in complex with glcnac-murnac (space group P1211) [Bacillus subtilis subsp. subtilis str. 168],4GYK_B Crystal structure of mutant (D318N) bacillus subtilis family 3 glycoside hydrolase (nagz) in complex with glcnac-murnac (space group P1211) [Bacillus subtilis subsp. subtilis str. 168] |
| 3SQL_A | 4.83e-48 | 61 | 359 | 31 | 335 | CrystalStructure of Glycoside Hydrolase from Synechococcus [Synechococcus sp. PCC 7002],3SQL_B Crystal Structure of Glycoside Hydrolase from Synechococcus [Synechococcus sp. PCC 7002],3SQM_A Crystal Structure of Glycoside Hydrolase from Synechococcus Complexed with N-acetyl-D-glucosamine [Synechococcus sp. PCC 7002],3SQM_B Crystal Structure of Glycoside Hydrolase from Synechococcus Complexed with N-acetyl-D-glucosamine [Synechococcus sp. PCC 7002],3SQM_C Crystal Structure of Glycoside Hydrolase from Synechococcus Complexed with N-acetyl-D-glucosamine [Synechococcus sp. PCC 7002],3SQM_D Crystal Structure of Glycoside Hydrolase from Synechococcus Complexed with N-acetyl-D-glucosamine [Synechococcus sp. PCC 7002] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P40406 | 2.03e-57 | 48 | 466 | 36 | 503 | Beta-hexosaminidase OS=Bacillus subtilis (strain 168) OX=224308 GN=nagZ PE=1 SV=1 |
| P48823 | 2.48e-46 | 80 | 430 | 55 | 435 | Beta-hexosaminidase A OS=Pseudoalteromonas piscicida OX=43662 GN=cht60 PE=1 SV=1 |
| B5RCU8 | 2.56e-43 | 581 | 984 | 30 | 406 | Putative D-alanyl-D-alanine carboxypeptidase OS=Salmonella gallinarum (strain 287/91 / NCTC 13346) OX=550538 GN=yfeW PE=3 SV=1 |
| B5BB26 | 4.68e-43 | 581 | 984 | 30 | 406 | Putative D-alanyl-D-alanine carboxypeptidase OS=Salmonella paratyphi A (strain AKU_12601) OX=554290 GN=yfeW PE=3 SV=1 |
| Q5PCQ2 | 4.68e-43 | 581 | 984 | 30 | 406 | Putative D-alanyl-D-alanine carboxypeptidase OS=Salmonella paratyphi A (strain ATCC 9150 / SARB42) OX=295319 GN=yfeW PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
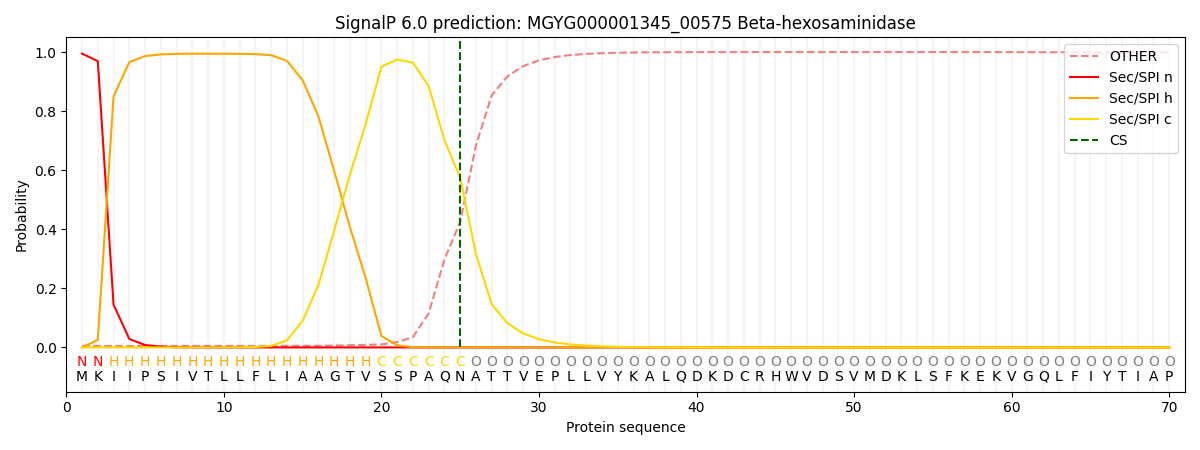
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.009868 | 0.988126 | 0.000758 | 0.000422 | 0.000370 | 0.000413 |
