You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001345_00624
Basic Information
help
| Species |
Bacteroides xylanisolvens
|
| Lineage |
Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Bacteroidaceae; Bacteroides; Bacteroides xylanisolvens
|
| CAZyme ID |
MGYG000001345_00624
|
| CAZy Family |
GH5 |
| CAZyme Description |
hypothetical protein
|
| CAZyme Property |
|
| Genome Property |
| Genome Assembly ID |
Genome Size |
Genome Type |
Country |
Continent |
| MGYG000001345 |
5913018 |
Isolate |
not provided |
not provided |
|
| Gene Location |
Start: 163704;
End: 164777
Strand: +
|
No EC number prediction in MGYG000001345_00624.
| Family |
Start |
End |
Evalue |
family coverage |
| GH5 |
53 |
315 |
3e-118 |
0.9963636363636363 |
| Cdd ID |
Domain |
E-Value |
qStart |
qEnd |
sStart |
sEnd |
Domain Description |
| pfam00150
|
Cellulase |
6.12e-05 |
145 |
306 |
87 |
266 |
Cellulase (glycosyl hydrolase family 5). |
This protein is predicted as LIPO
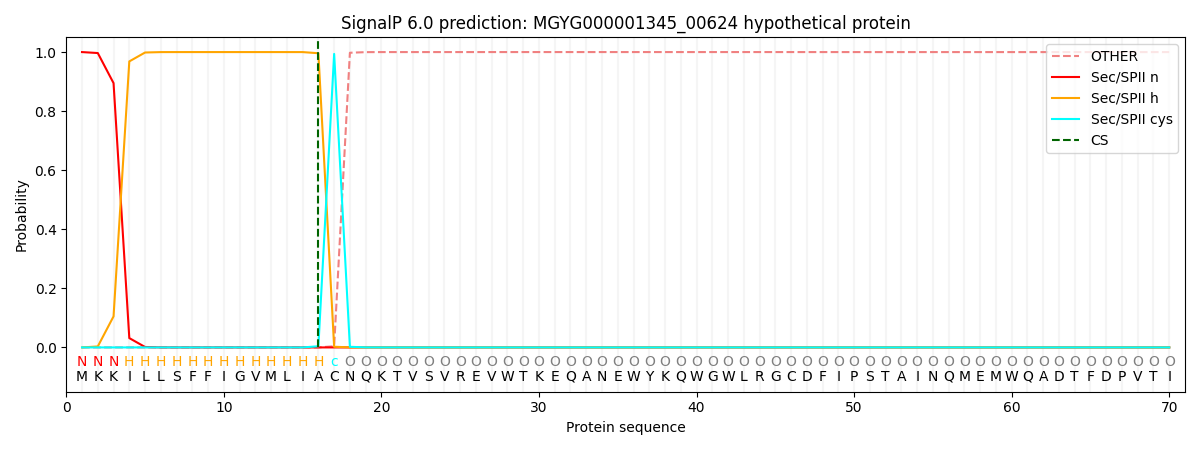
| Other |
SP_Sec_SPI |
LIPO_Sec_SPII |
TAT_Tat_SPI |
TATLIP_Sec_SPII |
PILIN_Sec_SPIII |
|
0.000000
|
0.000001
|
1.000078
|
0.000000
|
0.000000
|
0.000000
|
There is no transmembrane helices in MGYG000001345_00624.

