You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001345_01767
You are here: Home > Sequence: MGYG000001345_01767
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Bacteroides xylanisolvens | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Bacteroidaceae; Bacteroides; Bacteroides xylanisolvens | |||||||||||
| CAZyme ID | MGYG000001345_01767 | |||||||||||
| CAZy Family | PL13 | |||||||||||
| CAZyme Description | Heparin lyase I | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 133892; End: 135088 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| PL13 | 30 | 398 | 5.6e-208 | 0.9917355371900827 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam14099 | Polysacc_lyase | 2.80e-19 | 58 | 385 | 9 | 200 | Polysaccharide lyase. This family includes heparin lyase I, EC:4.2.2.7. Heparin lyase I depolymerizes heparin by cleaving the glycosidic linkage next to an iduronic acid moiety. The structure of heparin lyase I consists of a beta-jelly roll domain with a long, deep substrate-binding groove and an unusual thumb domain containing many basic residues extending from the main body of the enzyme. This family also includes glucuronan lyase, EC:4.2.2.14. The structure glucuronan lyase is a beta-jelly roll. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QUT22717.1 | 1.75e-298 | 1 | 398 | 1 | 398 |
| CBK68736.1 | 1.75e-298 | 1 | 398 | 1 | 398 |
| QUT28427.1 | 7.13e-298 | 1 | 398 | 1 | 398 |
| QQR18169.1 | 3.25e-294 | 1 | 398 | 1 | 398 |
| ANU56970.1 | 3.25e-294 | 1 | 398 | 1 | 398 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3ILR_A | 4.26e-241 | 31 | 398 | 3 | 370 | Structureof Heparinase I from Bacteroides thetaiotaomicron in complex with tetrasaccharide product [Bacteroides thetaiotaomicron] |
| 3IKW_A | 4.95e-241 | 31 | 398 | 7 | 374 | Structureof Heparinase I from Bacteroides thetaiotaomicron [Bacteroides thetaiotaomicron] |
| 3IMN_A | 3.32e-240 | 31 | 398 | 9 | 376 | Crystalstructure of heparin lyase I from Bacteroides thetaiotaomicron [Bacteroides thetaiotaomicron],3IN9_A Crystal structure of heparin lyase I complexed with disaccharide heparin [Bacteroides thetaiotaomicron] |
| 3INA_A | 1.11e-238 | 31 | 398 | 9 | 376 | Crystalstructure of heparin lyase I H151A mutant complexed with a dodecasaccharide heparin [Bacteroides thetaiotaomicron] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q05819 | 9.96e-173 | 1 | 397 | 1 | 381 | Heparin lyase I OS=Pedobacter heparinus OX=984 PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
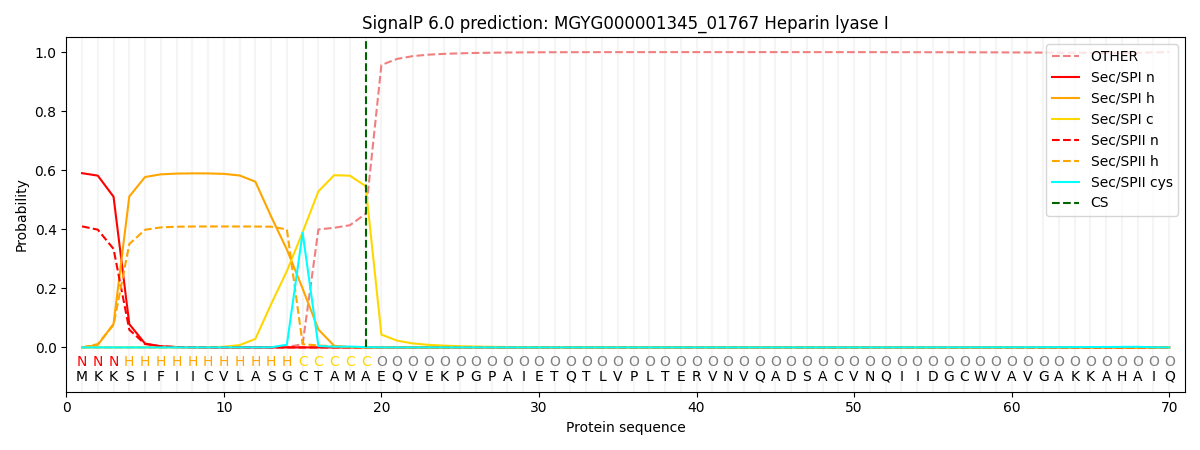
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000484 | 0.580474 | 0.418244 | 0.000346 | 0.000252 | 0.000200 |
