You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001345_03891
You are here: Home > Sequence: MGYG000001345_03891
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Bacteroides xylanisolvens | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Bacteroidaceae; Bacteroides; Bacteroides xylanisolvens | |||||||||||
| CAZyme ID | MGYG000001345_03891 | |||||||||||
| CAZy Family | GH18 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 471152; End: 472270 Strand: - | |||||||||||
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam16141 | DUF4849 | 6.28e-71 | 44 | 366 | 24 | 318 | Putative glycoside hydrolase Family 18, chitinase_18. This DUF is likely to be a form of glycosyl hydrolase from CAZy family 18, possibly chitinase 18. This would have the EC number of EC:3.2.1.14. |
| cd06542 | GH18_EndoS-like | 2.36e-53 | 57 | 347 | 1 | 242 | Endo-beta-N-acetylglucosaminidases are bacterial chitinases that hydrolyze the chitin core of various asparagine (N)-linked glycans and glycoproteins. The endo-beta-N-acetylglucosaminidases have a glycosyl hydrolase family 18 (GH18) catalytic domain. Some members also have an additional C-terminal glycosyl hydrolase family 20 (GH20) domain while others have an N-terminal domain of unknown function (pfam08522). Members of this family include endo-beta-N-acetylglucosaminidase S (EndoS) from Streptococcus pyogenes, EndoF1, EndoF2, EndoF3, and EndoH from Flavobacterium meningosepticum, and EndoE from Enterococcus faecalis. EndoS is a secreted endoglycosidase from Streptococcus pyogenes that specifically hydrolyzes the glycan on human IgG between two core N-acetylglucosamine residues. EndoE is a secreted endoglycosidase, encoded by the ndoE gene in Enterococcus faecalis, that hydrolyzes the glycan on human RNase B. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QRM99080.1 | 3.19e-290 | 1 | 372 | 1 | 372 |
| CBK67861.1 | 1.07e-288 | 1 | 372 | 1 | 372 |
| QUT32301.1 | 1.07e-288 | 1 | 372 | 1 | 372 |
| ALO49414.1 | 9.58e-166 | 1 | 371 | 1 | 365 |
| QUB44458.1 | 3.30e-163 | 1 | 372 | 1 | 363 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6Q64_A | 1.61e-60 | 3 | 368 | 2 | 356 | BT1044SeMetE190Q [Bacteroides thetaiotaomicron] |
| 6KPL_A | 6.56e-11 | 47 | 255 | 20 | 198 | CrystalStructure of endo-beta-N-acetylglucosaminidase from Cordyceps militaris in apo form [Cordyceps militaris CM01],6KPM_A Crystal Structure of endo-beta-N-acetylglucosaminidase from Cordyceps militaris in complex with L-fucose [Cordyceps militaris CM01] |
| 6KPN_A | 6.91e-10 | 47 | 255 | 20 | 198 | CrystalStructure of endo-beta-N-acetylglucosaminidase from Cordyceps militaris D154N/E156Q mutant in complex with fucosyl-N-acetylglucosamine [Cordyceps militaris CM01],6KPO_A Crystal Structure of endo-beta-N-acetylglucosaminidase from Cordyceps militaris D154N/E156Q mutant in complex with fucosyl-N-acetylglucosamine-Asn [Cordyceps militaris CM01] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P36912 | 5.01e-13 | 46 | 255 | 50 | 230 | Endo-beta-N-acetylglucosaminidase F2 OS=Elizabethkingia meningoseptica OX=238 GN=endOF2 PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as LIPO
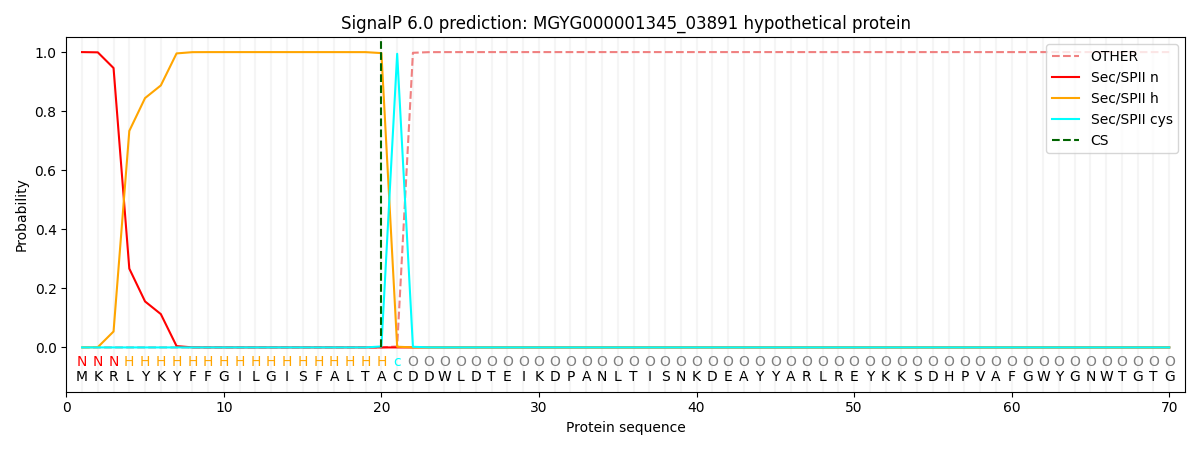
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000000 | 0.000000 | 1.000026 | 0.000000 | 0.000000 | 0.000000 |
