You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001354_01676
You are here: Home > Sequence: MGYG000001354_01676
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Bifidobacterium dentium | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Actinobacteriota; Actinomycetia; Actinomycetales; Bifidobacteriaceae; Bifidobacterium; Bifidobacterium dentium | |||||||||||
| CAZyme ID | MGYG000001354_01676 | |||||||||||
| CAZy Family | GH0 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 2056534; End: 2058537 Strand: - | |||||||||||
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG3408 | GDB1 | 4.03e-73 | 238 | 665 | 240 | 631 | Glycogen debranching enzyme (alpha-1,6-glucosidase) [Carbohydrate transport and metabolism]. |
| pfam06202 | GDE_C | 3.63e-14 | 260 | 631 | 9 | 367 | Amylo-alpha-1,6-glucosidase. This family includes human glycogen branching enzyme AGL. This enzyme contains a number of distinct catalytic activities. It has been shown for the yeast homolog GDB1 that mutations in this region disrupt the enzymes Amylo-alpha-1,6-glucosidase (EC:3.2.1.33). |
| pfam14742 | GDE_N_bis | 4.04e-11 | 20 | 213 | 1 | 192 | N-terminal domain of (some) glycogen debranching enzymes. This domain is found on the N-terminal of some glycogen debranching enzymes and is usually followed by the GDE_C (PF06202) and in this sense it is analogous (but probably not homologous) to the GDE_N (PF12439). Its exact function is unknown |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| VEG23875.1 | 0.0 | 1 | 667 | 1 | 667 |
| QTL78797.1 | 0.0 | 14 | 667 | 1 | 654 |
| BAQ27206.1 | 0.0 | 17 | 667 | 1 | 651 |
| ADB09890.1 | 0.0 | 52 | 667 | 1 | 616 |
| QAY32185.1 | 0.0 | 9 | 667 | 7 | 670 |
Swiss-Prot Hits help
SignalP and Lipop Annotations help
This protein is predicted as OTHER
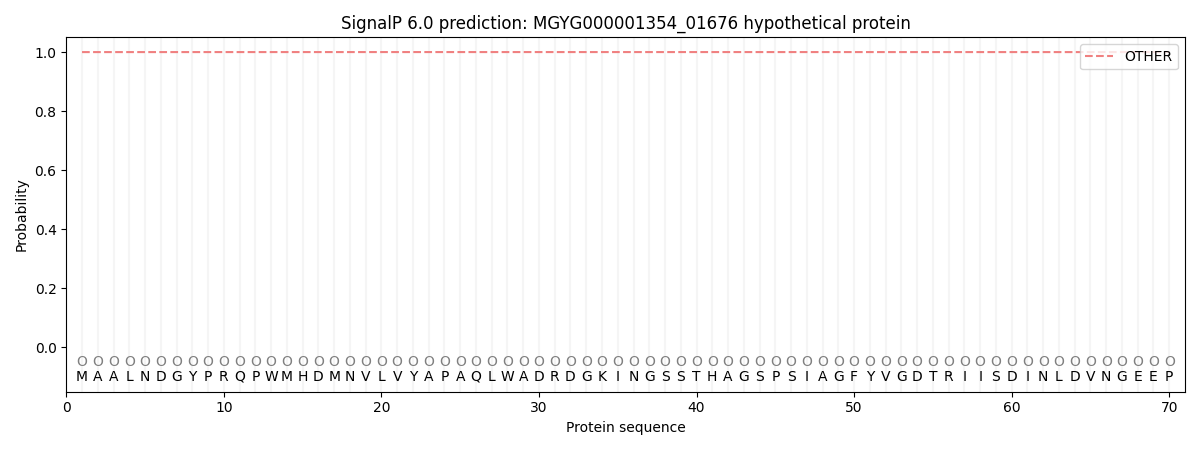
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000076 | 0.000001 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
