You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001364_03185
Basic Information
help
| Species |
Phocaeicola plebeius
|
| Lineage |
Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Bacteroidaceae; Phocaeicola; Phocaeicola plebeius
|
| CAZyme ID |
MGYG000001364_03185
|
| CAZy Family |
GH49 |
| CAZyme Description |
hypothetical protein
|
| CAZyme Property |
|
| Genome Property |
| Genome Assembly ID |
Genome Size |
Genome Type |
Country |
Continent |
| MGYG000001364 |
4421324 |
Isolate |
not provided |
Asia |
|
| Gene Location |
Start: 312446;
End: 314131
Strand: -
|
No EC number prediction in MGYG000001364_03185.
| Family |
Start |
End |
Evalue |
family coverage |
| GH49 |
53 |
472 |
5e-32 |
0.8269581056466302 |
MGYG000001364_03185 has no CDD domain.
This protein is predicted as LIPO
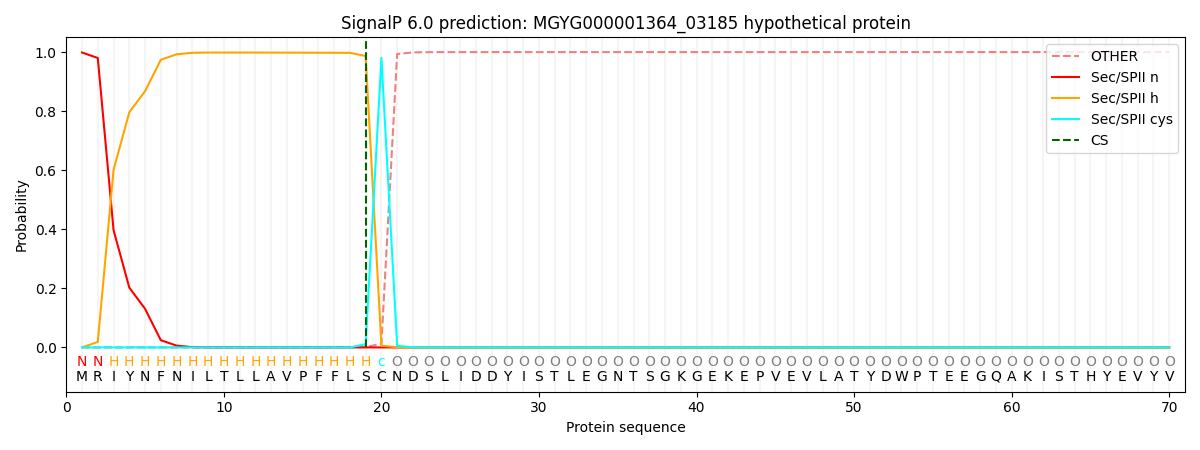
| Other |
SP_Sec_SPI |
LIPO_Sec_SPII |
TAT_Tat_SPI |
TATLIP_Sec_SPII |
PILIN_Sec_SPIII |
|
0.000015
|
0.001272
|
0.998736
|
0.000001
|
0.000001
|
0.000001
|
There is no transmembrane helices in MGYG000001364_03185.

