You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001371_00900
You are here: Home > Sequence: MGYG000001371_00900
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
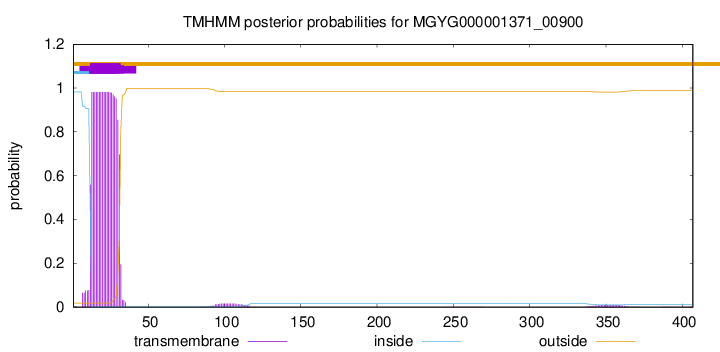
TMHMM annotations
Basic Information help
| Species | Paenibacillus lautus_A | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes; Bacilli; Paenibacillales; Paenibacillaceae; Paenibacillus; Paenibacillus lautus_A | |||||||||||
| CAZyme ID | MGYG000001371_00900 | |||||||||||
| CAZy Family | GH8 | |||||||||||
| CAZyme Description | Endoglucanase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 14692; End: 15915 Strand: - | |||||||||||
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam01270 | Glyco_hydro_8 | 1.68e-26 | 59 | 398 | 3 | 317 | Glycosyl hydrolases family 8. |
| COG3405 | BcsZ | 1.66e-08 | 64 | 171 | 29 | 132 | Endo-1,4-beta-D-glucanase Y [Carbohydrate transport and metabolism]. |
| PRK11097 | PRK11097 | 4.72e-04 | 90 | 171 | 52 | 133 | cellulase. |
| TIGR00920 | 2A060605 | 6.32e-04 | 229 | 348 | 644 | 755 | 3-hydroxy-3-methylglutaryl-coenzyme A reductase. [Transport and binding proteins, Carbohydrates, organic alcohols, and acids] |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| ACX62823.1 | 1.83e-300 | 1 | 407 | 1 | 407 |
| QOT08917.1 | 1.06e-299 | 1 | 407 | 1 | 407 |
| AYB46723.1 | 1.24e-298 | 1 | 407 | 1 | 407 |
| BAB69035.1 | 4.96e-289 | 1 | 407 | 1 | 407 |
| AVV55294.1 | 3.90e-286 | 1 | 407 | 1 | 407 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5XD0_A | 1.59e-227 | 7 | 407 | 8 | 409 | ApoStructure of Beta-1,3-1,4-glucanase from Paenibacillus sp.X4 [Paenibacillus sp. X4],5XD0_B Apo Structure of Beta-1,3-1,4-glucanase from Paenibacillus sp.X4 [Paenibacillus sp. X4] |
| 1V5C_A | 6.54e-98 | 33 | 405 | 5 | 382 | Thecrystal structure of the inactive form chitosanase from Bacillus sp. K17 at pH3.7 [Bacillus sp. (in: Bacteria)],1V5D_A The crystal structure of the active form chitosanase from Bacillus sp. K17 at pH6.4 [Bacillus sp. (in: Bacteria)],1V5D_B The crystal structure of the active form chitosanase from Bacillus sp. K17 at pH6.4 [Bacillus sp. (in: Bacteria)] |
| 7CJU_A | 8.92e-98 | 33 | 405 | 11 | 388 | Crystalstructure of inactive form of chitosanase crystallized by ammonium sulfate [Bacillus sp. K17-2],7CJU_B Crystal structure of inactive form of chitosanase crystallized by ammonium sulfate [Bacillus sp. K17-2],7XGQ_A Chain A, chitosanase [Bacillus sp. K17-2],7XGQ_B Chain B, chitosanase [Bacillus sp. K17-2] |
| 1CEM_A | 6.91e-47 | 31 | 403 | 1 | 355 | ChainA, CELLULASE CELA (1,4-BETA-D-GLUCAN-GLUCANOHYDROLASE) [Acetivibrio thermocellus],1IS9_A Chain A, endoglucanase A [Acetivibrio thermocellus] |
| 1KWF_A | 1.97e-45 | 31 | 403 | 1 | 355 | ChainA, Endoglucanase A [Acetivibrio thermocellus] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P19254 | 6.79e-224 | 11 | 407 | 12 | 409 | Beta-glucanase OS=Niallia circulans OX=1397 GN=bgc PE=3 SV=1 |
| P29019 | 9.16e-94 | 33 | 405 | 61 | 438 | Endoglucanase OS=Bacillus sp. (strain KSM-330) OX=72575 PE=1 SV=1 |
| A3DC29 | 3.98e-46 | 20 | 403 | 22 | 387 | Endoglucanase A OS=Acetivibrio thermocellus (strain ATCC 27405 / DSM 1237 / JCM 9322 / NBRC 103400 / NCIMB 10682 / NRRL B-4536 / VPI 7372) OX=203119 GN=celA PE=1 SV=1 |
| P37699 | 7.99e-43 | 14 | 403 | 16 | 388 | Endoglucanase C OS=Ruminiclostridium cellulolyticum (strain ATCC 35319 / DSM 5812 / JCM 6584 / H10) OX=394503 GN=celCCC PE=1 SV=2 |
| P37701 | 4.13e-42 | 18 | 403 | 20 | 388 | Endoglucanase 2 OS=Ruminiclostridium josui OX=1499 GN=celB PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP

| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000272 | 0.999070 | 0.000168 | 0.000163 | 0.000145 | 0.000134 |