You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001375_01375
You are here: Home > Sequence: MGYG000001375_01375
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Ruminococcus_F champanellensis | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Oscillospirales; Ruminococcaceae; Ruminococcus_F; Ruminococcus_F champanellensis | |||||||||||
| CAZyme ID | MGYG000001375_01375 | |||||||||||
| CAZy Family | PL1 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 1552887; End: 1555436 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| PL1 | 90 | 269 | 1.7e-54 | 0.9836065573770492 |
| CBM13 | 505 | 644 | 1.7e-21 | 0.6914893617021277 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam14200 | RicinB_lectin_2 | 5.08e-14 | 541 | 630 | 5 | 89 | Ricin-type beta-trefoil lectin domain-like. |
| cd14256 | Dockerin_I | 5.23e-14 | 797 | 849 | 2 | 56 | Type I dockerin repeat domain. Bacterial cohesin domains bind to a complementary protein domain named dockerin, and this interaction is required for the formation of the cellulosome, a cellulose-degrading complex. The cellulosome consists of scaffoldin, a noncatalytic scaffolding polypeptide, that comprises repeating cohesion modules and a single carbohydrate-binding module (CBM). Specific calcium-dependent interactions between cohesins and dockerins appear to be essential for cellulosome assembly. This subfamily represents type I dockerins, which are responsible for anchoring a variety of enzymatic domains to the complex. |
| pfam14200 | RicinB_lectin_2 | 5.28e-14 | 507 | 581 | 15 | 89 | Ricin-type beta-trefoil lectin domain-like. |
| pfam00404 | Dockerin_1 | 5.50e-08 | 797 | 849 | 1 | 55 | Dockerin type I repeat. The dockerin repeat is the binding partner of the cohesin domain pfam00963. The cohesin-dockerin interaction is the crucial interaction for complex formation in the cellulosome. The dockerin repeats, each bearing homology to the EF-hand calcium-binding loop bind calcium. |
| pfam14200 | RicinB_lectin_2 | 3.67e-07 | 587 | 644 | 1 | 56 | Ricin-type beta-trefoil lectin domain-like. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| CBL17517.1 | 0.0 | 1 | 849 | 1 | 849 |
| ADU22445.1 | 6.73e-276 | 1 | 789 | 1 | 728 |
| BAV13046.1 | 7.77e-216 | 41 | 490 | 50 | 510 |
| ADL50763.1 | 7.77e-216 | 41 | 490 | 50 | 510 |
| AUO19292.1 | 2.75e-210 | 23 | 788 | 16 | 644 |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| B8NQQ7 | 6.80e-49 | 39 | 481 | 20 | 418 | Probable pectate lyase C OS=Aspergillus flavus (strain ATCC 200026 / FGSC A1120 / IAM 13836 / NRRL 3357 / JCM 12722 / SRRC 167) OX=332952 GN=plyC PE=3 SV=1 |
| Q2UB83 | 1.13e-47 | 39 | 481 | 20 | 418 | Probable pectate lyase C OS=Aspergillus oryzae (strain ATCC 42149 / RIB 40) OX=510516 GN=plyC PE=3 SV=1 |
| Q5B297 | 5.07e-44 | 39 | 481 | 20 | 415 | Probable pectate lyase C OS=Emericella nidulans (strain FGSC A4 / ATCC 38163 / CBS 112.46 / NRRL 194 / M139) OX=227321 GN=plyC PE=3 SV=1 |
| Q4WL88 | 1.08e-42 | 39 | 481 | 21 | 419 | Probable pectate lyase C OS=Neosartorya fumigata (strain ATCC MYA-4609 / Af293 / CBS 101355 / FGSC A1100) OX=330879 GN=plyC PE=3 SV=1 |
| B0XMA2 | 1.46e-42 | 39 | 481 | 21 | 419 | Probable pectate lyase C OS=Neosartorya fumigata (strain CEA10 / CBS 144.89 / FGSC A1163) OX=451804 GN=plyC PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
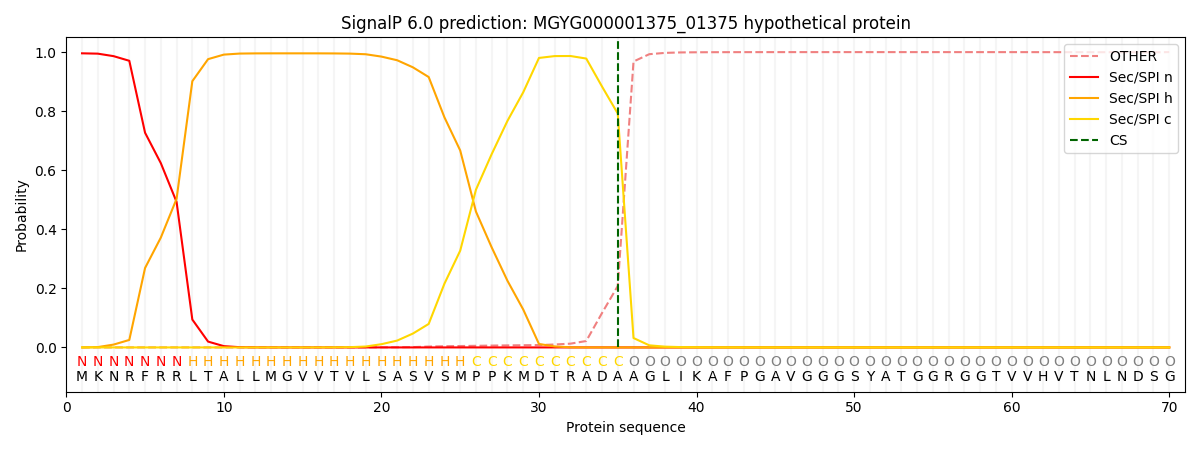
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000901 | 0.993225 | 0.004729 | 0.000657 | 0.000252 | 0.000203 |
