You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001388_00327
You are here: Home > Sequence: MGYG000001388_00327
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
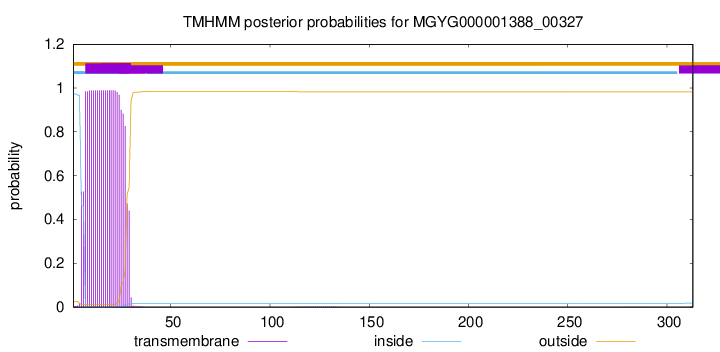
TMHMM annotations
Basic Information help
| Species | Desulfitobacterium hafniense | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_B; Desulfitobacteriia; Desulfitobacteriales; Desulfitobacteriaceae; Desulfitobacterium; Desulfitobacterium hafniense | |||||||||||
| CAZyme ID | MGYG000001388_00327 | |||||||||||
| CAZy Family | CBM2 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 39917; End: 40858 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| CBM2 | 192 | 295 | 6.5e-20 | 0.9207920792079208 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam00553 | CBM_2 | 8.91e-08 | 193 | 288 | 3 | 87 | Cellulose binding domain. Two tryptophan residues are involved in cellulose binding. Cellulose binding domain found in bacteria. |
| smart00637 | CBD_II | 2.85e-06 | 201 | 280 | 4 | 72 | CBD_II domain. |
| smart01063 | CBM49 | 1.15e-04 | 198 | 280 | 7 | 78 | Carbohydrate binding domain CBM49. This domain is found at the C terminal of cellulases and in vitro binding studies have shown it to binds to crystalline cellulose. |
| pfam09478 | CBM49 | 1.51e-04 | 206 | 280 | 14 | 76 | Carbohydrate binding domain CBM49. This domain is found at the C terminal of cellulases and in vitro binding studies have shown it to binds to crystalline cellulose. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| BAE86270.1 | 7.69e-225 | 1 | 313 | 1 | 313 |
| ACL18903.1 | 7.69e-225 | 1 | 313 | 1 | 313 |
| CDX04743.1 | 4.45e-224 | 1 | 313 | 1 | 313 |
| AFL99156.1 | 2.59e-194 | 1 | 313 | 1 | 316 |
| AFQ44073.1 | 2.56e-132 | 1 | 312 | 1 | 313 |
Swiss-Prot Hits help
SignalP and Lipop Annotations help
This protein is predicted as SP

| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.001677 | 0.997348 | 0.000279 | 0.000257 | 0.000210 | 0.000207 |