You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001388_02169
You are here: Home > Sequence: MGYG000001388_02169
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Desulfitobacterium hafniense | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_B; Desulfitobacteriia; Desulfitobacteriales; Desulfitobacteriaceae; Desulfitobacterium; Desulfitobacterium hafniense | |||||||||||
| CAZyme ID | MGYG000001388_02169 | |||||||||||
| CAZy Family | CBM50 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 102406; End: 103569 Strand: + | |||||||||||
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG3409 | PGRP | 3.48e-30 | 153 | 301 | 37 | 183 | Peptidoglycan-binding (PGRP) domain of peptidoglycan hydrolases [Cell wall/membrane/envelope biogenesis]. |
| COG3409 | PGRP | 5.51e-29 | 238 | 380 | 37 | 183 | Peptidoglycan-binding (PGRP) domain of peptidoglycan hydrolases [Cell wall/membrane/envelope biogenesis]. |
| COG3409 | PGRP | 4.04e-27 | 77 | 216 | 37 | 183 | Peptidoglycan-binding (PGRP) domain of peptidoglycan hydrolases [Cell wall/membrane/envelope biogenesis]. |
| pfam01471 | PG_binding_1 | 2.06e-20 | 245 | 301 | 1 | 57 | Putative peptidoglycan binding domain. This domain is composed of three alpha helices. This domain is found at the N or C-terminus of a variety of enzymes involved in bacterial cell wall degradation. This domain may have a general peptidoglycan binding function. This family is found N-terminal to the catalytic domain of matrixins. The domain is found to bind peptidoglycan experimentally. |
| pfam01471 | PG_binding_1 | 4.97e-19 | 324 | 380 | 1 | 57 | Putative peptidoglycan binding domain. This domain is composed of three alpha helices. This domain is found at the N or C-terminus of a variety of enzymes involved in bacterial cell wall degradation. This domain may have a general peptidoglycan binding function. This family is found N-terminal to the catalytic domain of matrixins. The domain is found to bind peptidoglycan experimentally. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| CDX01843.1 | 8.27e-274 | 1 | 387 | 1 | 390 |
| BAE83569.1 | 1.40e-273 | 1 | 387 | 15 | 404 |
| ACL20957.1 | 4.08e-272 | 1 | 387 | 7 | 391 |
| AGA69465.1 | 7.92e-226 | 1 | 385 | 7 | 380 |
| AFM00719.1 | 4.83e-175 | 1 | 305 | 7 | 299 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5NM7_A | 9.34e-11 | 318 | 385 | 4 | 70 | Crystalstructure of Burkholderia AP3 phage endolysin [Burkholderia],5NM7_G Crystal structure of Burkholderia AP3 phage endolysin [Burkholderia] |
| 3BKH_A | 3.13e-10 | 240 | 339 | 12 | 110 | ChainA, lytic transglycosylase [Pseudomonas phage phiKZ],3BKV_A Chain A, lytic transglycosylase [Pseudomonas phage phiKZ] |
| 5TV7_A | 3.62e-09 | 319 | 381 | 109 | 170 | ChainA, Putative peptidoglycan-binding/hydrolysing protein [Clostridioides difficile 630],5TV7_B Chain B, Putative peptidoglycan-binding/hydrolysing protein [Clostridioides difficile 630] |
| 6TCI_A | 5.53e-08 | 78 | 136 | 6 | 64 | Thecrystal structure of SleB N-terminal domain [Bacillus cereus ATCC 14579] |
| 4LPQ_A | 1.14e-07 | 241 | 301 | 12 | 72 | Crystalstructure of the L,D-transpeptidase (residues 123-326) from Xylanimonas cellulosilytica DSM 15894 [Xylanimonas cellulosilytica DSM 15894] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q99125 | 9.52e-10 | 240 | 376 | 195 | 345 | Probable N-acetylmuramoyl-L-alanine amidase OS=Bacillus licheniformis OX=1402 PE=3 SV=1 |
| P36550 | 1.31e-09 | 240 | 376 | 196 | 351 | N-acetylmuramoyl-L-alanine amidase CwlL OS=Bacillus licheniformis OX=1402 GN=cwlL PE=3 SV=1 |
| O31447 | 1.76e-09 | 167 | 380 | 84 | 276 | Uncharacterized protein YbfG OS=Bacillus subtilis (strain 168) OX=224308 GN=ybfG PE=3 SV=1 |
| L7N653 | 4.78e-08 | 155 | 301 | 12 | 160 | N-acetylmuramoyl-L-alanine amidase CwlM OS=Mycobacterium tuberculosis (strain ATCC 25618 / H37Rv) OX=83332 GN=cwlM PE=1 SV=1 |
| P49320 | 1.82e-07 | 238 | 301 | 41 | 105 | Uncharacterized protein in bpoA1 3'region (Fragment) OS=Kitasatospora aureofaciens OX=1894 PE=4 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
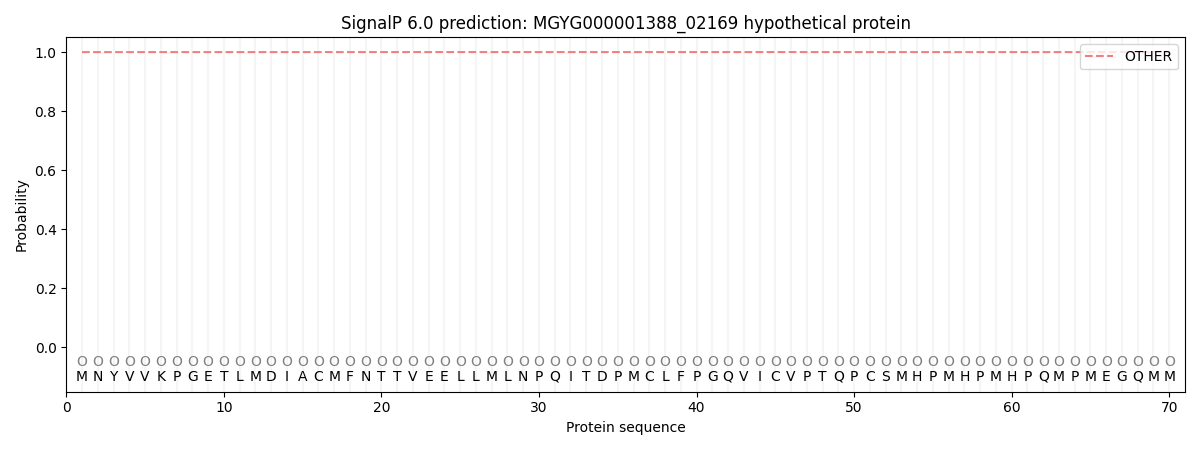
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000043 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
