You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001391_02538
You are here: Home > Sequence: MGYG000001391_02538
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Coprobacter fastidiosus | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Coprobacteraceae; Coprobacter; Coprobacter fastidiosus | |||||||||||
| CAZyme ID | MGYG000001391_02538 | |||||||||||
| CAZy Family | GH33 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 133815; End: 136100 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH33 | 200 | 532 | 5.9e-85 | 0.9415204678362573 |
| CE3 | 573 | 742 | 3.5e-16 | 0.8041237113402062 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd15482 | Sialidase_non-viral | 4.85e-102 | 201 | 550 | 2 | 339 | Non-viral sialidases. Sialidases or neuraminidases function to bind and hydrolyze terminal sialic acid residues from various glycoconjugates, they play vital roles in pathogenesis, bacterial nutrition and cellular interactions. They have a six-bladed, beta-propeller fold with the non-viral sialidases containing 2-5 Asp-box motifs (most commonly Ser/Thr-X-Asp-[X]-Gly-X-Thr- Trp/Phe). This CD includes eubacterial and eukaryotic sialidases. |
| cd01828 | sialate_O-acetylesterase_like2 | 6.45e-56 | 586 | 760 | 1 | 169 | sialate_O-acetylesterase_like subfamily of the SGNH-hydrolases, a diverse family of lipases and esterases. The tertiary fold of the enzyme is substantially different from that of the alpha/beta hydrolase family and unique among all known hydrolases; its active site closely resembles the Ser-His-Asp(Glu) triad found in other serine hydrolases. |
| cd01841 | NnaC_like | 6.98e-34 | 585 | 759 | 1 | 174 | NnaC (CMP-NeuNAc synthetase) _like subfamily of SGNH_hydrolases, a diverse family of lipases and esterases. The tertiary fold of the enzyme is substantially different from that of the alpha/beta hydrolase family and unique among all known hydrolases; its active site closely resembles two of the three components of typical Ser-His-Asp(Glu) triad from other serine hydrolases. E. coli NnaC appears to be involved in polysaccharide synthesis. |
| pfam13472 | Lipase_GDSL_2 | 1.80e-24 | 589 | 743 | 1 | 169 | GDSL-like Lipase/Acylhydrolase family. This family of presumed lipases and related enzymes are similar to pfam00657. |
| pfam13088 | BNR_2 | 7.07e-23 | 243 | 525 | 15 | 280 | BNR repeat-like domain. This family of proteins contains BNR-like repeats suggesting these proteins may act as sialidases. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QCX54140.1 | 1.59e-127 | 65 | 567 | 44 | 534 |
| BBD46557.1 | 5.73e-124 | 65 | 559 | 9 | 493 |
| SCM56345.1 | 7.49e-117 | 65 | 555 | 49 | 529 |
| QKJ63384.1 | 9.39e-113 | 122 | 561 | 545 | 969 |
| BBL05903.1 | 9.40e-112 | 202 | 559 | 32 | 382 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6XPM_A | 1.34e-82 | 562 | 760 | 3 | 202 | ChainA, Lysophospholipase L1 [Phocaeicola vulgatus] |
| 6NJC_A | 7.23e-82 | 562 | 760 | 3 | 202 | ChainA, Sialate O-acetylesterase [Phocaeicola vulgatus ATCC 8482],6NJC_B Chain B, Sialate O-acetylesterase [Phocaeicola vulgatus ATCC 8482],6XPG_A Chain A, Lysophospholipase L1 [Phocaeicola vulgatus] |
| 6MNJ_A | 2.61e-45 | 75 | 528 | 43 | 521 | Hadzamicrobial sialidase Hz136 [Alistipes],6MNJ_B Hadza microbial sialidase Hz136 [Alistipes] |
| 1EUR_A | 2.55e-41 | 206 | 530 | 18 | 346 | Sialidase[Micromonospora viridifaciens],1EUS_A Sialidase Complexed With 2-Deoxy-2,3-Dehydro-N- Acetylneuraminic Acid [Micromonospora viridifaciens] |
| 6MYV_A | 4.45e-41 | 86 | 528 | 33 | 504 | Sialidase26co-crystallized with DANA-Gc [bacterium],6MYV_B Sialidase26 co-crystallized with DANA-Gc [bacterium],6MYV_C Sialidase26 co-crystallized with DANA-Gc [bacterium],6MYV_D Sialidase26 co-crystallized with DANA-Gc [bacterium] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q02834 | 6.41e-39 | 206 | 564 | 60 | 411 | Sialidase OS=Micromonospora viridifaciens OX=1881 GN=nedA PE=1 SV=1 |
| P31206 | 1.32e-37 | 81 | 528 | 50 | 526 | Sialidase OS=Bacteroides fragilis (strain YCH46) OX=295405 GN=nanH PE=3 SV=2 |
| P29767 | 1.59e-20 | 202 | 554 | 384 | 826 | Sialidase OS=Clostridium septicum OX=1504 PE=3 SV=1 |
| P15698 | 3.24e-16 | 210 | 440 | 51 | 282 | Sialidase OS=Paeniclostridium sordellii OX=1505 PE=1 SV=1 |
| P10481 | 6.55e-14 | 210 | 495 | 33 | 324 | Sialidase OS=Clostridium perfringens OX=1502 GN=nanH PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
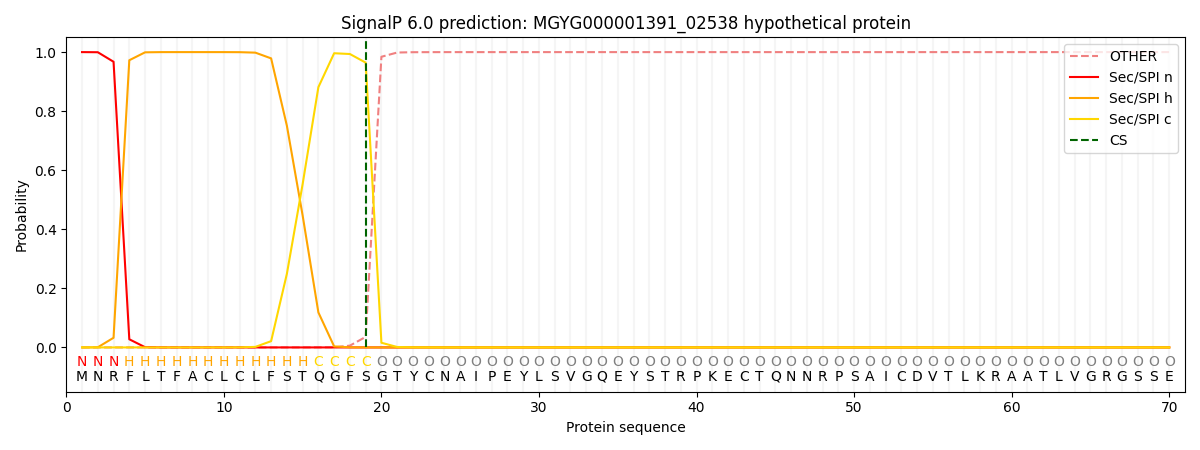
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000305 | 0.999058 | 0.000201 | 0.000141 | 0.000137 | 0.000134 |
