You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001399_02110
You are here: Home > Sequence: MGYG000001399_02110
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Odoribacter laneus | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Marinifilaceae; Odoribacter; Odoribacter laneus | |||||||||||
| CAZyme ID | MGYG000001399_02110 | |||||||||||
| CAZy Family | GT1 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 2512663; End: 2513805 Strand: + | |||||||||||
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam13528 | Glyco_trans_1_3 | 1.59e-22 | 1 | 323 | 1 | 309 | Glycosyl transferase family 1. |
| COG1819 | YjiC | 2.35e-05 | 1 | 315 | 2 | 346 | UDP:flavonoid glycosyltransferase YjiC, YdhE family [Carbohydrate transport and metabolism]. |
| cd03784 | GT1_Gtf-like | 1.37e-04 | 1 | 324 | 1 | 362 | UDP-glycosyltransferases and similar proteins. This family includes the Gtfs, a group of homologous glycosyltransferases involved in the final stages of the biosynthesis of antibiotics vancomycin and related chloroeremomycin. Gtfs transfer sugar moieties from an activated NDP-sugar donor to the oxidatively cross-linked heptapeptide core of vancomycin group antibiotics. The core structure is important for the bioactivity of the antibiotics. |
| COG0707 | MurG | 0.001 | 1 | 322 | 1 | 309 | UDP-N-acetylglucosamine:LPS N-acetylglucosamine transferase [Cell wall/membrane/envelope biogenesis]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| BCI61991.1 | 1.27e-150 | 1 | 355 | 1 | 355 |
| QUT49491.1 | 1.68e-136 | 1 | 345 | 1 | 345 |
| QCY55033.1 | 5.51e-135 | 1 | 345 | 1 | 345 |
| QUT52939.1 | 5.51e-135 | 1 | 345 | 1 | 345 |
| QIX65697.1 | 5.51e-135 | 1 | 345 | 1 | 345 |
Swiss-Prot Hits help
SignalP and Lipop Annotations help
This protein is predicted as OTHER
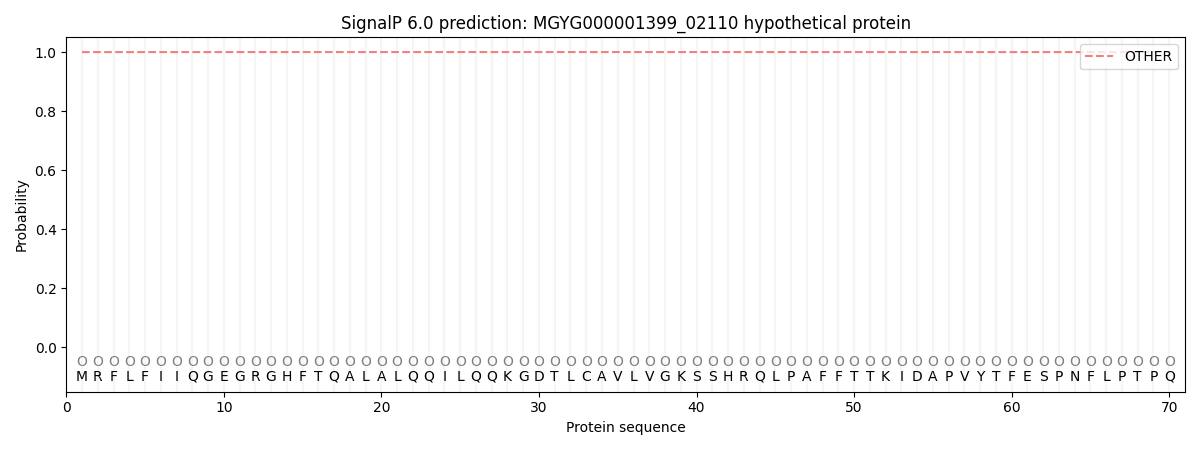
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000057 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
