You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001400_00790
You are here: Home > Sequence: MGYG000001400_00790
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
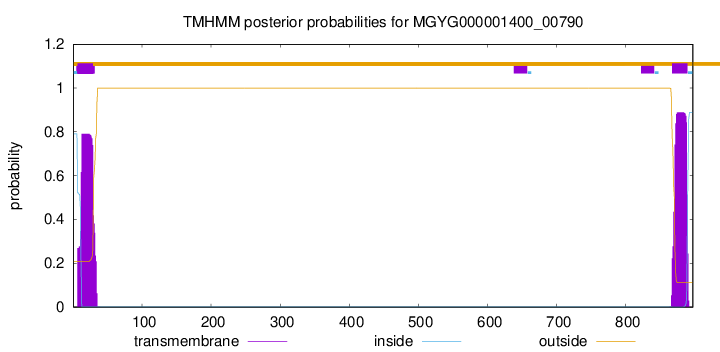
TMHMM annotations
Basic Information help
| Species | Erysipelatoclostridium ramosum | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes; Bacilli; Erysipelotrichales; Erysipelatoclostridiaceae; Erysipelatoclostridium; Erysipelatoclostridium ramosum | |||||||||||
| CAZyme ID | MGYG000001400_00790 | |||||||||||
| CAZy Family | GH3 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 831569; End: 834262 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH3 | 228 | 479 | 1.1e-42 | 0.8796296296296297 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG1472 | BglX | 6.34e-32 | 153 | 571 | 7 | 352 | Periplasmic beta-glucosidase and related glycosidases [Carbohydrate transport and metabolism]. |
| pfam00933 | Glyco_hydro_3 | 2.27e-25 | 247 | 479 | 89 | 281 | Glycosyl hydrolase family 3 N terminal domain. |
| PRK15098 | PRK15098 | 1.87e-14 | 247 | 788 | 121 | 600 | beta-glucosidase BglX. |
| pfam01915 | Glyco_hydro_3_C | 5.53e-07 | 722 | 805 | 122 | 207 | Glycosyl hydrolase family 3 C-terminal domain. This domain is involved in catalysis and may be involved in binding beta-glucan. This domain is found associated with pfam00933. |
| PLN03080 | PLN03080 | 0.003 | 235 | 438 | 102 | 297 | Probable beta-xylosidase; Provisional |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QQV06598.1 | 0.0 | 1 | 897 | 1 | 897 |
| QPS11706.1 | 0.0 | 1 | 897 | 1 | 897 |
| QMW75023.1 | 0.0 | 1 | 897 | 1 | 897 |
| QQY26495.1 | 0.0 | 1 | 897 | 1 | 897 |
| AQR97785.1 | 7.72e-294 | 66 | 836 | 47 | 799 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5M6G_A | 2.63e-16 | 246 | 575 | 143 | 430 | Crystalstructure Glucan 1,4-beta-glucosidase from Saccharopolyspora erythraea [Saccharopolyspora erythraea D] |
| 5TF0_A | 1.29e-15 | 246 | 794 | 99 | 591 | CrystalStructure of Glycosil Hydrolase Family 3 N-Terminal Domain Protein from Bacteroides intestinalis [Bacteroides intestinalis DSM 17393],5TF0_B Crystal Structure of Glycosil Hydrolase Family 3 N-Terminal Domain Protein from Bacteroides intestinalis [Bacteroides intestinalis DSM 17393] |
| 5XXL_A | 2.26e-15 | 244 | 794 | 98 | 592 | Crystalstructure of GH3 beta-glucosidase from Bacteroides thetaiotaomicron [Bacteroides thetaiotaomicron VPI-5482],5XXL_B Crystal structure of GH3 beta-glucosidase from Bacteroides thetaiotaomicron [Bacteroides thetaiotaomicron VPI-5482],5XXM_A Crystal structure of GH3 beta-glucosidase from Bacteroides thetaiotaomicron in complex with gluconolactone [Bacteroides thetaiotaomicron VPI-5482],5XXM_B Crystal structure of GH3 beta-glucosidase from Bacteroides thetaiotaomicron in complex with gluconolactone [Bacteroides thetaiotaomicron VPI-5482] |
| 4ZOA_A | 6.57e-15 | 177 | 648 | 49 | 468 | CrystalStructure of beta-glucosidase from Listeria innocua in complex with isofagomine [Listeria innocua Clip11262],4ZOA_B Crystal Structure of beta-glucosidase from Listeria innocua in complex with isofagomine [Listeria innocua Clip11262],4ZOB_A Crystal Structure of beta-glucosidase from Listeria innocua in complex with gluconolactone [Listeria innocua Clip11262],4ZOB_B Crystal Structure of beta-glucosidase from Listeria innocua in complex with gluconolactone [Listeria innocua Clip11262],4ZOD_A Crystal Structure of beta-glucosidase from Listeria innocua in complex with glucose [Listeria innocua Clip11262],4ZOD_B Crystal Structure of beta-glucosidase from Listeria innocua in complex with glucose [Listeria innocua Clip11262],4ZOE_A Crystal Structure of beta-glucosidase from Listeria innocua [Listeria innocua Clip11262],4ZOE_B Crystal Structure of beta-glucosidase from Listeria innocua [Listeria innocua Clip11262] |
| 5XXN_A | 8.92e-15 | 244 | 794 | 98 | 592 | CrystalStructure of mutant (D286N) beta-glucosidase from Bacteroides thetaiotaomicron in complex with sophorose [Bacteroides thetaiotaomicron VPI-5482],5XXN_B Crystal Structure of mutant (D286N) beta-glucosidase from Bacteroides thetaiotaomicron in complex with sophorose [Bacteroides thetaiotaomicron VPI-5482],5XXO_A Crystal structure of mutant (D286N) GH3 beta-glucosidase from Bacteroides thetaiotaomicron in complex with sophorotriose [Bacteroides thetaiotaomicron VPI-5482],5XXO_B Crystal structure of mutant (D286N) GH3 beta-glucosidase from Bacteroides thetaiotaomicron in complex with sophorotriose [Bacteroides thetaiotaomicron VPI-5482] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q46684 | 2.47e-61 | 111 | 562 | 46 | 486 | Periplasmic beta-glucosidase/beta-xylosidase OS=Dickeya chrysanthemi OX=556 GN=bgxA PE=3 SV=1 |
| Q2UFP8 | 1.59e-57 | 195 | 825 | 119 | 627 | Probable beta-glucosidase C OS=Aspergillus oryzae (strain ATCC 42149 / RIB 40) OX=510516 GN=bglC PE=3 SV=2 |
| Q5BCC6 | 7.21e-57 | 196 | 824 | 108 | 615 | Beta-glucosidase C OS=Emericella nidulans (strain FGSC A4 / ATCC 38163 / CBS 112.46 / NRRL 194 / M139) OX=227321 GN=bglC PE=1 SV=1 |
| B8NGU6 | 2.42e-56 | 195 | 825 | 115 | 623 | Probable beta-glucosidase C OS=Aspergillus flavus (strain ATCC 200026 / FGSC A1120 / IAM 13836 / NRRL 3357 / JCM 12722 / SRRC 167) OX=332952 GN=bglC PE=3 SV=1 |
| Q23892 | 9.97e-15 | 240 | 567 | 184 | 480 | Lysosomal beta glucosidase OS=Dictyostelium discoideum OX=44689 GN=gluA PE=1 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as SP
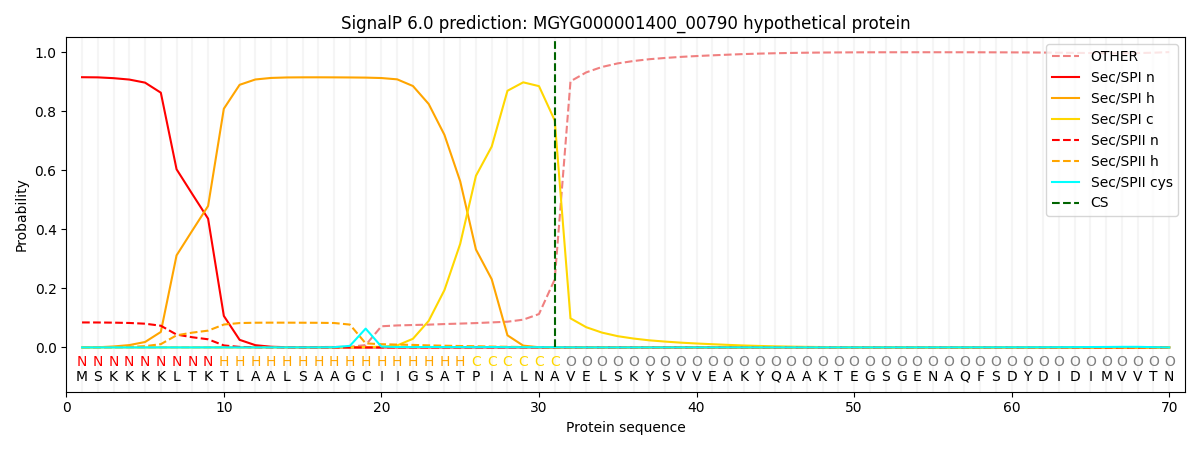
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.002149 | 0.903198 | 0.093168 | 0.000676 | 0.000419 | 0.000342 |