You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001400_02791
You are here: Home > Sequence: MGYG000001400_02791
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
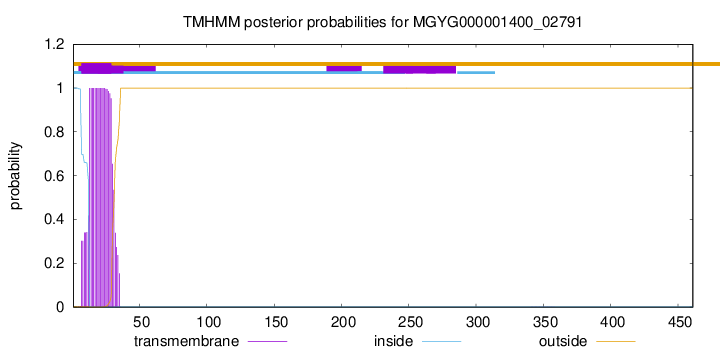
TMHMM annotations
Basic Information help
| Species | Erysipelatoclostridium ramosum | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes; Bacilli; Erysipelotrichales; Erysipelatoclostridiaceae; Erysipelatoclostridium; Erysipelatoclostridium ramosum | |||||||||||
| CAZyme ID | MGYG000001400_02791 | |||||||||||
| CAZy Family | GH3 | |||||||||||
| CAZyme Description | Beta-hexosaminidase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 1190554; End: 1191939 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH3 | 195 | 419 | 8.6e-41 | 0.9768518518518519 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam00933 | Glyco_hydro_3 | 4.81e-56 | 142 | 453 | 1 | 315 | Glycosyl hydrolase family 3 N terminal domain. |
| COG1472 | BglX | 2.46e-51 | 141 | 461 | 1 | 318 | Periplasmic beta-glucosidase and related glycosidases [Carbohydrate transport and metabolism]. |
| PRK05337 | PRK05337 | 3.63e-45 | 161 | 430 | 18 | 290 | beta-hexosaminidase; Provisional |
| PRK15098 | PRK15098 | 9.27e-18 | 134 | 460 | 38 | 358 | beta-glucosidase BglX. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QQV05565.1 | 0.0 | 1 | 461 | 1 | 461 |
| QQY27485.1 | 0.0 | 1 | 461 | 1 | 461 |
| QMW75963.1 | 0.0 | 1 | 461 | 1 | 461 |
| QPS13697.1 | 0.0 | 1 | 461 | 1 | 461 |
| BCI61427.1 | 2.86e-120 | 46 | 461 | 63 | 482 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6K5J_A | 2.19e-32 | 141 | 460 | 11 | 342 | Structureof a glycoside hydrolase family 3 beta-N-acetylglucosaminidase from Paenibacillus sp. str. FPU-7 [Paenibacillaceae] |
| 5BU9_A | 6.47e-28 | 141 | 455 | 5 | 337 | Crystalstructure of Beta-N-acetylhexosaminidase from Beutenbergia cavernae DSM 12333 [Beutenbergia cavernae DSM 12333],5BU9_B Crystal structure of Beta-N-acetylhexosaminidase from Beutenbergia cavernae DSM 12333 [Beutenbergia cavernae DSM 12333] |
| 6GFV_A | 7.58e-26 | 146 | 455 | 21 | 337 | Mtuberculosis LpqI [Mycobacterium tuberculosis H37Rv],6GFV_B M tuberculosis LpqI [Mycobacterium tuberculosis H37Rv] |
| 4ZM6_A | 3.15e-25 | 143 | 457 | 9 | 339 | Aunique GCN5-related glucosamine N-acetyltransferase region exist in the fungal multi-domain GH3 beta-N-acetylglucosaminidase [Rhizomucor miehei CAU432],4ZM6_B A unique GCN5-related glucosamine N-acetyltransferase region exist in the fungal multi-domain GH3 beta-N-acetylglucosaminidase [Rhizomucor miehei CAU432] |
| 2OXN_A | 4.70e-25 | 162 | 425 | 19 | 277 | Vibriocholerae family 3 glycoside hydrolase (NagZ) in complex with PUGNAc [Vibrio cholerae] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q4USG7 | 6.00e-33 | 158 | 449 | 13 | 309 | Beta-hexosaminidase OS=Xanthomonas campestris pv. campestris (strain 8004) OX=314565 GN=nagZ PE=3 SV=1 |
| Q8PB42 | 6.00e-33 | 158 | 449 | 13 | 309 | Beta-hexosaminidase OS=Xanthomonas campestris pv. campestris (strain ATCC 33913 / DSM 3586 / NCPPB 528 / LMG 568 / P 25) OX=190485 GN=nagZ PE=3 SV=1 |
| B0RX17 | 6.00e-33 | 158 | 449 | 13 | 309 | Beta-hexosaminidase OS=Xanthomonas campestris pv. campestris (strain B100) OX=509169 GN=nagZ PE=3 SV=1 |
| B2I6G9 | 6.24e-32 | 158 | 449 | 13 | 309 | Beta-hexosaminidase OS=Xylella fastidiosa (strain M23) OX=405441 GN=nagZ PE=3 SV=1 |
| Q9PAZ0 | 6.24e-32 | 158 | 449 | 13 | 309 | Beta-hexosaminidase OS=Xylella fastidiosa (strain 9a5c) OX=160492 GN=nagZ PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
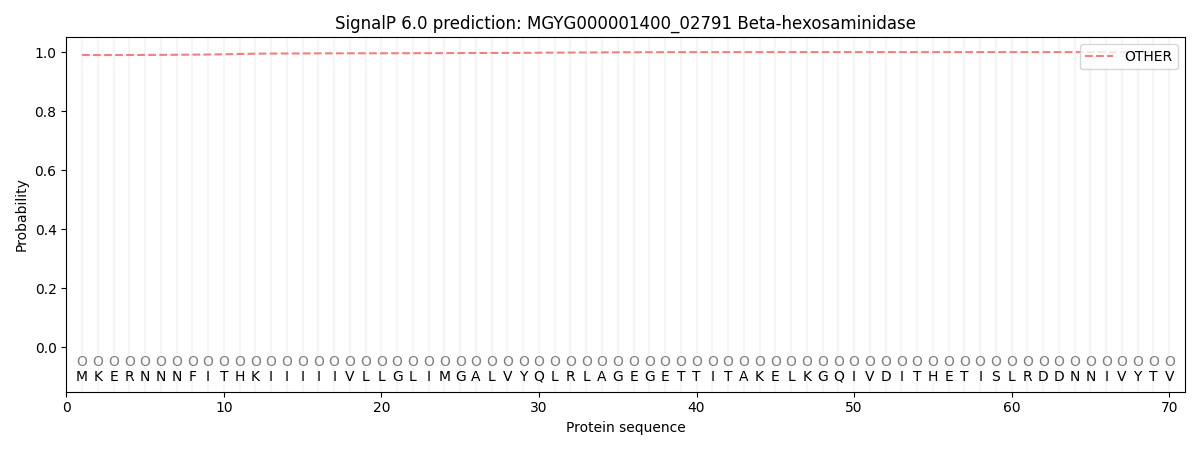
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.990257 | 0.004226 | 0.000319 | 0.000024 | 0.000016 | 0.005180 |