You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001407_02476
You are here: Home > Sequence: MGYG000001407_02476
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Clostridium_J senegalense | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Clostridiales; Clostridiaceae; Clostridium_J; Clostridium_J senegalense | |||||||||||
| CAZyme ID | MGYG000001407_02476 | |||||||||||
| CAZy Family | GH73 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 233631; End: 234527 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH73 | 148 | 287 | 1.6e-33 | 0.9765625 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG1705 | FlgJ | 1.12e-44 | 136 | 291 | 40 | 188 | Flagellum-specific peptidoglycan hydrolase FlgJ [Cell wall/membrane/envelope biogenesis, Cell motility]. |
| NF038016 | sporang_Gsm | 2.98e-24 | 123 | 287 | 145 | 308 | sporangiospore maturation cell wall hydrolase GsmA. The peptidoglycan-hydrolyzing enzyme GsmA occurs in some sporangia-forming members of the Actinobacteria, such as Actinoplanes missouriensis, and is required for proper separation of spores. GsmA proteins have one or two SH3 domains N-terminal to the hydrolase domain. |
| PRK05684 | flgJ | 4.84e-23 | 141 | 283 | 154 | 295 | flagellar assembly peptidoglycan hydrolase FlgJ. |
| smart00047 | LYZ2 | 7.70e-22 | 141 | 289 | 10 | 143 | Lysozyme subfamily 2. Eubacterial enzymes distantly related to eukaryotic lysozymes. |
| pfam01832 | Glucosaminidase | 1.01e-21 | 147 | 229 | 1 | 77 | Mannosyl-glycoprotein endo-beta-N-acetylglucosaminidase. This family includes Mannosyl-glycoprotein endo-beta-N-acetylglucosaminidase EC:3.2.1.96. As well as the flageller protein J that has been shown to hydrolyze peptidoglycan. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| CEJ72889.1 | 8.39e-97 | 2 | 297 | 4 | 296 |
| QYE98805.1 | 5.51e-95 | 2 | 297 | 4 | 296 |
| AXU64913.1 | 1.62e-94 | 2 | 296 | 4 | 295 |
| AXU50442.1 | 1.62e-94 | 2 | 296 | 4 | 295 |
| QQY61939.1 | 3.26e-94 | 2 | 294 | 4 | 293 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3FI7_A | 1.76e-17 | 139 | 291 | 30 | 183 | CrystalStructure of the autolysin Auto (Lmo1076) from Listeria monocytogenes, catalytic domain [Listeria monocytogenes EGD-e] |
| 5T1Q_A | 1.24e-09 | 134 | 293 | 57 | 214 | ChainA, N-acetylmuramoyl-L-alanine amidase domain-containing protein SAOUHSC_02979 [Staphylococcus aureus subsp. aureus NCTC 8325],5T1Q_B Chain B, N-acetylmuramoyl-L-alanine amidase domain-containing protein SAOUHSC_02979 [Staphylococcus aureus subsp. aureus NCTC 8325],5T1Q_C Chain C, N-acetylmuramoyl-L-alanine amidase domain-containing protein SAOUHSC_02979 [Staphylococcus aureus subsp. aureus NCTC 8325],5T1Q_D Chain D, N-acetylmuramoyl-L-alanine amidase domain-containing protein SAOUHSC_02979 [Staphylococcus aureus subsp. aureus NCTC 8325] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| O32083 | 3.92e-25 | 130 | 291 | 35 | 198 | Exo-glucosaminidase LytG OS=Bacillus subtilis (strain 168) OX=224308 GN=lytG PE=1 SV=1 |
| Q9X9J3 | 8.67e-15 | 134 | 281 | 155 | 301 | Peptidoglycan hydrolase FlgJ OS=Vibrio parahaemolyticus serotype O3:K6 (strain RIMD 2210633) OX=223926 GN=flgJ PE=3 SV=1 |
| Q9CIT4 | 1.35e-10 | 142 | 291 | 65 | 214 | Probable N-acetylmuramidase OS=Lactococcus lactis subsp. lactis (strain IL1403) OX=272623 GN=acmA PE=3 SV=1 |
| Q9KQ15 | 1.77e-10 | 141 | 265 | 163 | 292 | Peptidoglycan hydrolase FlgJ OS=Vibrio cholerae serotype O1 (strain ATCC 39315 / El Tor Inaba N16961) OX=243277 GN=flgJ PE=3 SV=2 |
| A2RHZ5 | 4.35e-10 | 142 | 291 | 65 | 214 | Probable N-acetylmuramidase OS=Lactococcus lactis subsp. cremoris (strain MG1363) OX=416870 GN=acmA PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
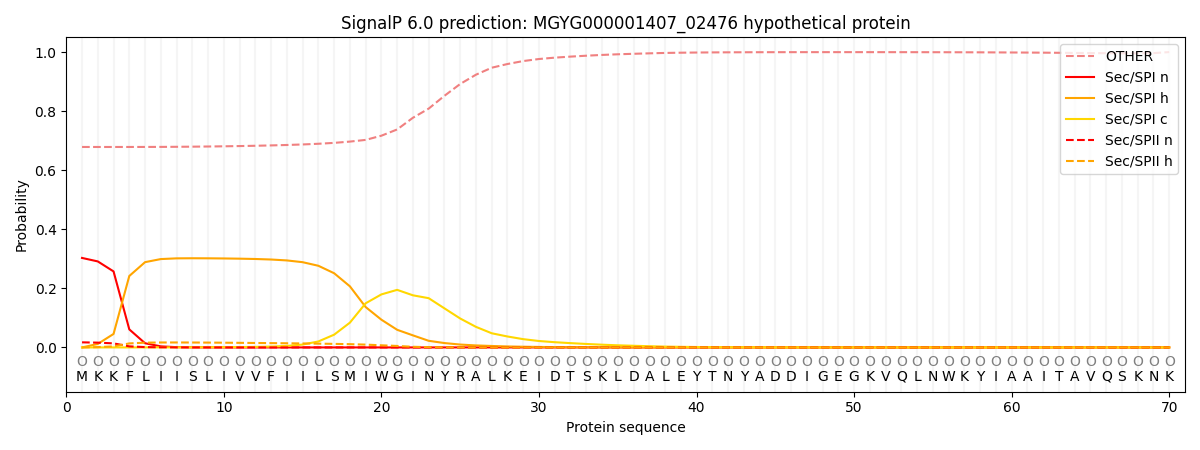
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.693382 | 0.284434 | 0.018917 | 0.000764 | 0.000458 | 0.002044 |

