You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001418_02636
You are here: Home > Sequence: MGYG000001418_02636
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
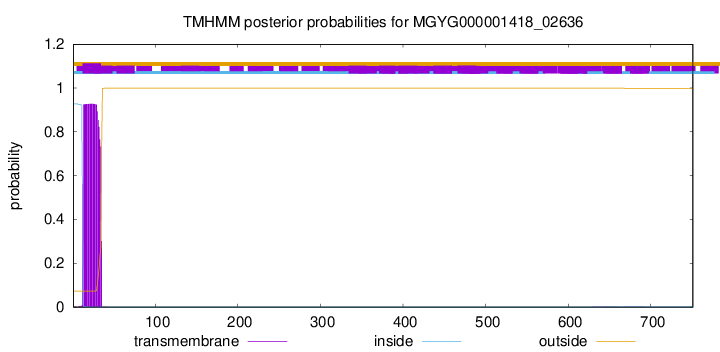
TMHMM annotations
Basic Information help
| Species | Aeromicrobium massiliense | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Actinobacteriota; Actinomycetia; Propionibacteriales; Nocardioidaceae; Aeromicrobium; Aeromicrobium massiliense | |||||||||||
| CAZyme ID | MGYG000001418_02636 | |||||||||||
| CAZy Family | CBM5 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 99408; End: 101663 Strand: - | |||||||||||
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| PRK09419 | PRK09419 | 1.01e-80 | 36 | 557 | 657 | 1137 | multifunctional 2',3'-cyclic-nucleotide 2'-phosphodiesterase/3'-nucleotidase/5'-nucleotidase. |
| COG0737 | UshA | 6.23e-77 | 31 | 555 | 18 | 501 | 2',3'-cyclic-nucleotide 2'-phosphodiesterase/5'- or 3'-nucleotidase, 5'-nucleotidase family [Nucleotide transport and metabolism, Defense mechanisms]. |
| PRK09558 | ushA | 3.84e-61 | 28 | 558 | 22 | 540 | bifunctional UDP-sugar hydrolase/5'-nucleotidase periplasmic precursor; Reviewed |
| cd07412 | MPP_YhcR_N | 4.04e-55 | 40 | 312 | 1 | 294 | Bacillus subtilis YhcR endonuclease and related proteins, N-terminal metallophosphatase domain. YhcR is a Bacillus subtilis sugar-nonspecific endonuclease. It cleaves endonucleolytically to yield nucleotide 3'-monophosphate products, similar to Staphylococcus aureus micrococcal nuclease. YhcR appears to be located in the cell wall, and is thought to be a substrate for a Bacillus subtilis sortase. YhcR is the major calcium-activated nuclease of B. subtilis. The N-terminal metallophosphatase domain belongs to a large superfamily of distantly related metallophosphatases (MPPs) that includes: Mre11/SbcD-like exonucleases, Dbr1-like RNA lariat debranching enzymes, YfcE-like phosphodiesterases, purple acid phosphatases (PAPs), YbbF-like UDP-2,3-diacylglucosamine hydrolases, and acid sphingomyelinases (ASMases). MPPs are functionally diverse, but all share a conserved domain with an active site consisting of two metal ions (usually manganese, iron, or zinc) coordinated with octahedral geometry by a cage of histidine, aspartate, and asparagine residues. The conserved domain is a double beta-sheet sandwich with a di-metal active site made up of residues located at the C-terminal side of the sheets. This domain is thought to allow for productive metal coordination. |
| pfam02872 | 5_nucleotid_C | 6.13e-41 | 358 | 529 | 4 | 153 | 5'-nucleotidase, C-terminal domain. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QAY64865.1 | 3.91e-220 | 27 | 629 | 845 | 1448 |
| QAY71724.1 | 6.14e-215 | 27 | 611 | 838 | 1429 |
| ACZ32093.1 | 2.34e-211 | 26 | 629 | 823 | 1440 |
| SDS92078.1 | 5.53e-206 | 32 | 629 | 830 | 1437 |
| QAY73698.1 | 1.92e-188 | 38 | 629 | 906 | 1494 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1OI8_A | 4.33e-31 | 40 | 558 | 9 | 514 | 5'-Nucleotidase(E. coli) with an Engineered Disulfide Bridge (P90C, L424C) [Escherichia coli],1OI8_B 5'-Nucleotidase (E. coli) with an Engineered Disulfide Bridge (P90C, L424C) [Escherichia coli] |
| 1HO5_A | 7.24e-31 | 40 | 558 | 9 | 514 | 5'-Nucleotidase(E. Coli) In Complex With Adenosine And Phosphate [Escherichia coli],1HO5_B 5'-Nucleotidase (E. Coli) In Complex With Adenosine And Phosphate [Escherichia coli],1HPU_A 5'-Nucleotidase (Closed Form), Complex With Ampcp [Escherichia coli],1HPU_B 5'-Nucleotidase (Closed Form), Complex With Ampcp [Escherichia coli],1HPU_C 5'-Nucleotidase (Closed Form), Complex With Ampcp [Escherichia coli],1HPU_D 5'-Nucleotidase (Closed Form), Complex With Ampcp [Escherichia coli] |
| 1USH_A | 9.23e-31 | 40 | 558 | 34 | 539 | 5'-NucleotidaseFrom E. Coli [Escherichia coli K-12],2USH_A 5'-Nucleotidase From E. Coli [Escherichia coli K-12],2USH_B 5'-Nucleotidase From E. Coli [Escherichia coli K-12] |
| 1HP1_A | 2.13e-30 | 40 | 558 | 9 | 505 | 5'-Nucleotidase(Open Form) Complex With Atp [Escherichia coli] |
| 1OID_A | 2.50e-30 | 40 | 558 | 9 | 514 | 5'-Nucleotidase(E. coli) with an Engineered Disulfide Bridge (S228C, P513C) [Escherichia coli],1OID_B 5'-Nucleotidase (E. coli) with an Engineered Disulfide Bridge (S228C, P513C) [Escherichia coli],1OIE_A 5'-Nucleotidase (E. coli) with an Engineered Disulfide Bridge (S228C, P513C) [Escherichia coli] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q8YAJ5 | 1.40e-41 | 20 | 550 | 14 | 563 | Cell wall protein Lmo0130 OS=Listeria monocytogenes serovar 1/2a (strain ATCC BAA-679 / EGD-e) OX=169963 GN=lmo0130 PE=1 SV=1 |
| P54602 | 1.54e-41 | 36 | 556 | 586 | 1076 | Endonuclease YhcR OS=Bacillus subtilis (strain 168) OX=224308 GN=yhcR PE=1 SV=1 |
| Q56878 | 1.29e-34 | 40 | 558 | 34 | 539 | Protein UshA OS=Yersinia enterocolitica serotype O:8 / biotype 1B (strain NCTC 13174 / 8081) OX=393305 GN=ushA PE=3 SV=3 |
| P07024 | 5.05e-30 | 40 | 558 | 34 | 539 | Protein UshA OS=Escherichia coli (strain K12) OX=83333 GN=ushA PE=1 SV=2 |
| Q8DFG4 | 9.50e-29 | 19 | 555 | 10 | 539 | 5'-nucleotidase OS=Vibrio vulnificus (strain CMCP6) OX=216895 GN=nutA PE=3 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as SP
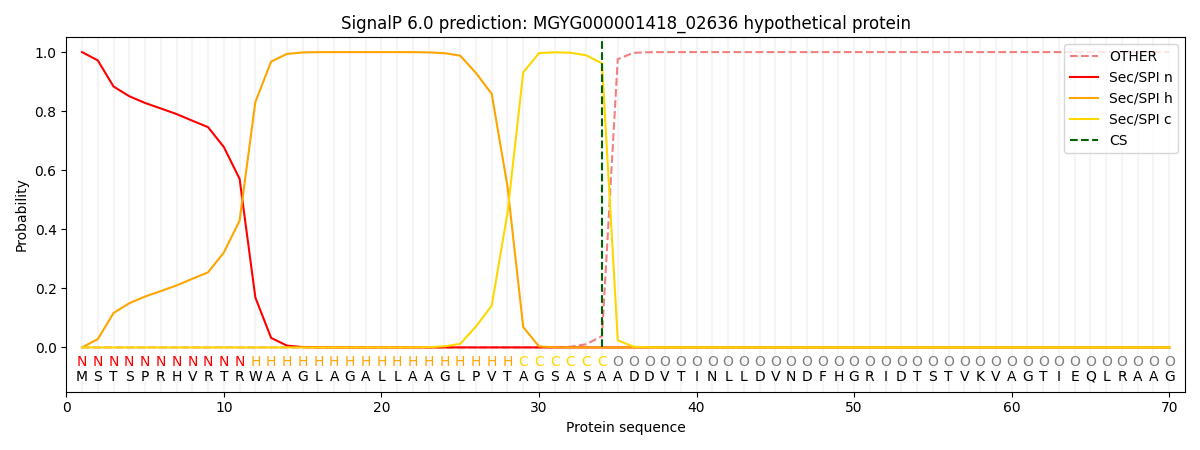
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000314 | 0.998956 | 0.000183 | 0.000235 | 0.000171 | 0.000148 |