You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001420_03141
Basic Information
help
| Species |
Alistipes senegalensis
|
| Lineage |
Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Rikenellaceae; Alistipes; Alistipes senegalensis
|
| CAZyme ID |
MGYG000001420_03141
|
| CAZy Family |
GH5 |
| CAZyme Description |
hypothetical protein
|
| CAZyme Property |
|
| Genome Property |
| Genome Assembly ID |
Genome Size |
Genome Type |
Country |
Continent |
| MGYG000001420 |
3985129 |
Isolate |
not provided |
not provided |
|
| Gene Location |
Start: 1715819;
End: 1716925
Strand: +
|
No EC number prediction in MGYG000001420_03141.
| Family |
Start |
End |
Evalue |
family coverage |
| GH5 |
54 |
316 |
4.2e-92 |
0.9927272727272727 |
| Cdd ID |
Domain |
E-Value |
qStart |
qEnd |
sStart |
sEnd |
Domain Description |
| COG3934
|
COG3934 |
7.82e-06 |
41 |
195 |
2 |
148 |
Endo-1,4-beta-mannosidase [Carbohydrate transport and metabolism]. |
This protein is predicted as LIPO
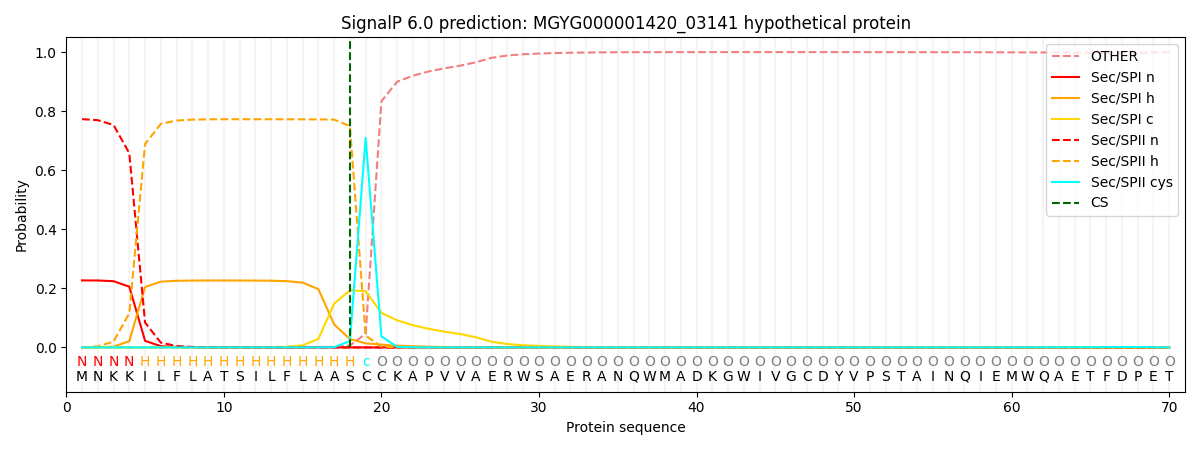
| Other |
SP_Sec_SPI |
LIPO_Sec_SPII |
TAT_Tat_SPI |
TATLIP_Sec_SPII |
PILIN_Sec_SPIII |
|
0.000864
|
0.219469
|
0.778891
|
0.000292
|
0.000281
|
0.000212
|
There is no transmembrane helices in MGYG000001420_03141.

