You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001422_01848
You are here: Home > Sequence: MGYG000001422_01848
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Bacteroides oleiciplenus | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Bacteroidaceae; Bacteroides; Bacteroides oleiciplenus | |||||||||||
| CAZyme ID | MGYG000001422_01848 | |||||||||||
| CAZy Family | GH10 | |||||||||||
| CAZyme Description | Endo-1,4-beta-xylanase/feruloyl esterase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 174093; End: 175187 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH10 | 30 | 362 | 8.7e-113 | 0.9867986798679867 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam00331 | Glyco_hydro_10 | 2.08e-124 | 30 | 363 | 1 | 310 | Glycosyl hydrolase family 10. |
| smart00633 | Glyco_10 | 1.31e-116 | 71 | 361 | 1 | 263 | Glycosyl hydrolase family 10. |
| COG3693 | XynA | 3.89e-103 | 10 | 364 | 6 | 340 | Endo-1,4-beta-xylanase, GH35 family [Carbohydrate transport and metabolism]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| ALJ59290.1 | 4.10e-210 | 12 | 364 | 567 | 919 |
| QDO67768.1 | 8.21e-210 | 12 | 364 | 567 | 919 |
| QUT89652.1 | 9.35e-209 | 12 | 364 | 567 | 919 |
| QUT46563.1 | 1.87e-137 | 1 | 364 | 1 | 360 |
| QRQ47387.1 | 1.87e-137 | 1 | 364 | 1 | 360 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5AY7_A | 1.57e-134 | 25 | 364 | 14 | 346 | Apsychrophilic glycoside hydrolase family 10 endo-beta-1,4-xylanase [Aegilops speltoides subsp. speltoides],5AY7_B A psychrophilic glycoside hydrolase family 10 endo-beta-1,4-xylanase [Aegilops speltoides subsp. speltoides],5D4Y_A A psychrophilic glycoside hydrolase family 10 endo-beta-1,4-xylanase [environmental samples],5D4Y_B A psychrophilic glycoside hydrolase family 10 endo-beta-1,4-xylanase [environmental samples] |
| 6WQW_A | 3.76e-105 | 30 | 362 | 6 | 327 | ChainA, Beta-xylanase [Thermobacillus composti] |
| 3EMC_A | 1.16e-100 | 29 | 362 | 6 | 328 | Crystalstructure of XynB, an intracellular xylanase from Paenibacillus barcinonensis [Paenibacillus barcinonensis],3EMQ_A Crystal structure of xilanase XynB from Paenibacillus barcelonensis complexed with an inhibitor [Paenibacillus barcinonensis],3EMZ_A Crystal structure of xylanase XynB from Paenibacillus barcinonensis complexed with a conduramine derivative [Paenibacillus barcinonensis] |
| 1N82_A | 2.14e-98 | 29 | 363 | 7 | 330 | Thehigh-resolution crystal structure of IXT6, a thermophilic, intracellular xylanase from G. stearothermophilus [Geobacillus stearothermophilus],1N82_B The high-resolution crystal structure of IXT6, a thermophilic, intracellular xylanase from G. stearothermophilus [Geobacillus stearothermophilus],3MUA_A Enzyme-Substrate interactions of IXT6, the intracellular xylanase of G. stearothermophilus. [Geobacillus stearothermophilus],3MUA_B Enzyme-Substrate interactions of IXT6, the intracellular xylanase of G. stearothermophilus. [Geobacillus stearothermophilus] |
| 2Q8X_A | 1.22e-97 | 29 | 363 | 7 | 330 | Thehigh-resolution crystal structure of ixt6, a thermophilic, intracellular xylanase from G. stearothermophilus [unidentified],2Q8X_B The high-resolution crystal structure of ixt6, a thermophilic, intracellular xylanase from G. stearothermophilus [unidentified],3MSD_A Enzyme-Substrate interactions of IXT6, the intracellular xylanase of G. stearothermophilus. [Geobacillus stearothermophilus],3MSD_B Enzyme-Substrate interactions of IXT6, the intracellular xylanase of G. stearothermophilus. [Geobacillus stearothermophilus],3MSG_A Enzyme-Substrate interactions of IXT6, the intracellular xylanase of G. stearothermophilus. [Geobacillus stearothermophilus],3MSG_B Enzyme-Substrate interactions of IXT6, the intracellular xylanase of G. stearothermophilus. [Geobacillus stearothermophilus] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| D5EY13 | 6.56e-121 | 1 | 364 | 1 | 369 | Endo-1,4-beta-xylanase/feruloyl esterase OS=Prevotella ruminicola (strain ATCC 19189 / JCM 8958 / 23) OX=264731 GN=xyn10D-fae1A PE=1 SV=1 |
| O69231 | 6.58e-100 | 29 | 362 | 7 | 329 | Endo-1,4-beta-xylanase B OS=Paenibacillus barcinonensis OX=198119 GN=xynB PE=1 SV=1 |
| P45703 | 2.38e-94 | 29 | 363 | 7 | 329 | Endo-1,4-beta-xylanase OS=Geobacillus stearothermophilus OX=1422 GN=xynA PE=1 SV=1 |
| P10474 | 6.59e-84 | 2 | 362 | 3 | 370 | Endoglucanase/exoglucanase B OS=Caldicellulosiruptor saccharolyticus OX=44001 GN=celB PE=3 SV=1 |
| Q12603 | 1.17e-83 | 11 | 364 | 13 | 352 | Beta-1,4-xylanase OS=Dictyoglomus thermophilum OX=14 GN=xynA PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as LIPO
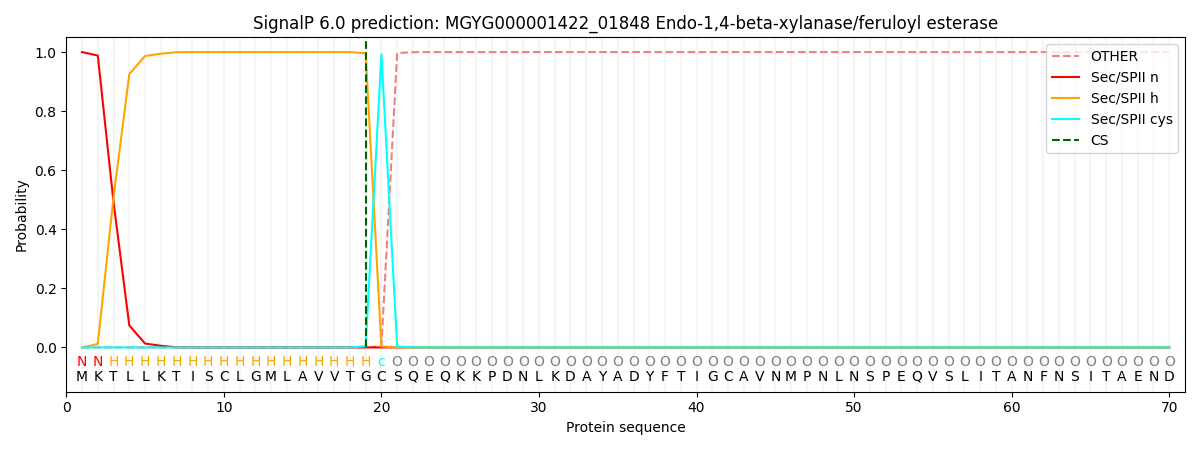
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000000 | 0.000000 | 1.000059 | 0.000000 | 0.000000 | 0.000000 |
