You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001423_00519
You are here: Home > Sequence: MGYG000001423_00519
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
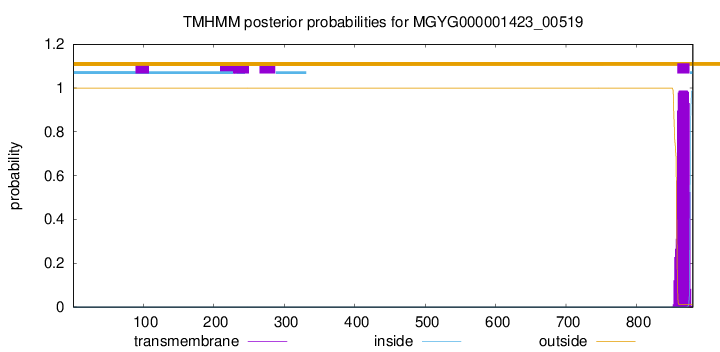
TMHMM annotations
Basic Information help
| Species | Clostridium celatum | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Clostridiales; Clostridiaceae; Clostridium; Clostridium celatum | |||||||||||
| CAZyme ID | MGYG000001423_00519 | |||||||||||
| CAZy Family | GH101 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 47163; End: 49805 Strand: - | |||||||||||
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam17451 | Glyco_hyd_101C | 1.01e-14 | 8 | 55 | 58 | 111 | Glycosyl hydrolase 101 beta sandwich domain. Virulence of pathogenic organisms such as the Gram-positive Streptococcus pneumoniae is largely determined by the ability to degrade host glycoproteins and to metabolize the resultant carbohydrates. This family is the enzymatic region, EC:3.2.1.97, of the cell surface proteins that specifically cleave Gal-beta-1,3-GalNAc-alpha-Ser/Thr (T-antigen, galacto-N-biose), the core 1 type O-linked glycan common to mucin glycoproteins. This reaction is exemplified by a S. pneumoniae protein, where Asp764 is the catalytic nucleophile-base and Glu796 the catalytic proton donor. This domain represents C-terminal the beta sandwich domain. |
| pfam17974 | GalBD_like | 1.44e-12 | 241 | 342 | 1 | 114 | Galactose-binding domain-like. Proteins containing a galactose-binding domain-like fold can be found in several different protein families, in both eukaryotes and prokaryotes. The common function of these domains is to bind to specific ligands, such as cell-surface-attached carbohydrate substrates for galactose oxidase and sialidase, phospholipids on the outer side of the mammalian cell membrane for coagulation factor Va, membrane-anchored ephrin for the Eph family of receptor tyrosine kinases, and a complex of broken single-stranded DNA and DNA polymerase beta for XRCC1. The structure of the galactose-binding domain-like members consists of a beta-sandwich, in which the strands making up the sheets exhibit a jellyroll fold. |
| pfam07554 | FIVAR | 2.77e-07 | 546 | 592 | 20 | 69 | FIVAR domain. This domain is found in a wide variety of contexts, but mostly occurring in cell wall associated proteins. A lack of conserved catalytic residues suggests that it is a binding domain. From context, possible substrates are hyaluronate or fibronectin (personal obs: C Yeats). This is further evidenced by. Possibly the exact substrate is N-acetyl glucosamine. Finding it in the same protein as pfam05089 further supports this proposal. It is found in the C-terminal part of Bacillus sp. Gellan lyase, which is removed during maturation. Some of the proteins it is found in are involved in methicillin resistance. The name FIVAR derives from Found In Various Architectures. |
| pfam02018 | CBM_4_9 | 5.83e-04 | 95 | 230 | 1 | 129 | Carbohydrate binding domain. This family includes diverse carbohydrate binding domains. |
| TIGR01167 | LPXTG_anchor | 7.61e-04 | 849 | 880 | 1 | 32 | LPXTG-motif cell wall anchor domain. This model describes the LPXTG motif-containing region found at the C-terminus of many surface proteins of Streptococcus and Streptomyces species. Cleavage between the Thr and Gly by sortase or a related enzyme leads to covalent anchoring at the new C-terminal Thr to the cell wall. Hits that do not lie at the C-terminus or are not found in Gram-positive bacteria are probably false-positive. A common feature of this proteins containing this domain appears to be a high proportion of charged and zwitterionic residues immediatedly upstream of the LPXTG motif. This model differs from other descriptions of the LPXTG region by including a portion of that upstream charged region. [Cell envelope, Other] |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QAS61722.1 | 6.80e-279 | 1 | 880 | 1076 | 1929 |
| AYE33559.1 | 6.80e-279 | 1 | 880 | 1076 | 1929 |
| ATD54516.1 | 1.14e-253 | 1 | 880 | 1073 | 1927 |
| ATD57802.1 | 1.14e-253 | 1 | 880 | 1073 | 1927 |
| QBJ74801.1 | 1.14e-253 | 1 | 880 | 1073 | 1927 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5A55_A | 3.22e-32 | 10 | 342 | 654 | 1007 | Thenative structure of GH101 from Streptococcus pneumoniae TIGR4 [Streptococcus pneumoniae TIGR4] |
| 5A56_A | 3.23e-32 | 10 | 342 | 652 | 1005 | Thestructure of GH101 from Streptococcus pneumoniae TIGR4 in complex with 1-O-methyl-T-antigen [Streptococcus pneumoniae TIGR4] |
| 5A58_A | 3.23e-32 | 10 | 342 | 652 | 1005 | Thestructure of GH101 D764N mutant from Streptococcus pneumoniae TIGR4 in complex with serinyl T-antigen [Streptococcus pneumoniae TIGR4] |
| 5A57_A | 3.23e-32 | 10 | 342 | 652 | 1005 | Thestructure of GH101 from Streptococcus pneumoniae TIGR4 in complex with PUGT [Streptococcus pneumoniae TIGR4] |
| 5A59_A | 3.23e-32 | 10 | 342 | 652 | 1005 | Thestructure of GH101 E796Q mutant from Streptococcus pneumoniae TIGR4 in complex with T-antigen [Streptococcus pneumoniae TIGR4],5A5A_A The structure of GH101 E796Q mutant from Streptococcus pneumoniae TIGR4 in complex with PNP-T-antigen [Streptococcus pneumoniae TIGR4] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q2MGH6 | 2.45e-31 | 10 | 342 | 967 | 1320 | Endo-alpha-N-acetylgalactosaminidase OS=Streptococcus pneumoniae serotype 4 (strain ATCC BAA-334 / TIGR4) OX=170187 GN=SP_0368 PE=1 SV=1 |
| Q8DR60 | 5.60e-31 | 10 | 342 | 967 | 1320 | Endo-alpha-N-acetylgalactosaminidase OS=Streptococcus pneumoniae (strain ATCC BAA-255 / R6) OX=171101 GN=spr0328 PE=1 SV=1 |
| A9WNA0 | 8.08e-30 | 13 | 443 | 643 | 1061 | Putative endo-alpha-N-acetylgalactosaminidase OS=Renibacterium salmoninarum (strain ATCC 33209 / DSM 20767 / JCM 11484 / NBRC 15589 / NCIMB 2235) OX=288705 GN=RSal33209_1326 PE=3 SV=2 |
| P29767 | 1.04e-09 | 415 | 589 | 842 | 1014 | Sialidase OS=Clostridium septicum OX=1504 PE=3 SV=1 |
| E8MGH9 | 1.22e-09 | 523 | 733 | 1661 | 1872 | Beta-L-arabinobiosidase OS=Bifidobacterium longum subsp. longum (strain ATCC 15707 / DSM 20219 / JCM 1217 / NCTC 11818 / E194b) OX=565042 GN=hypBA2 PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
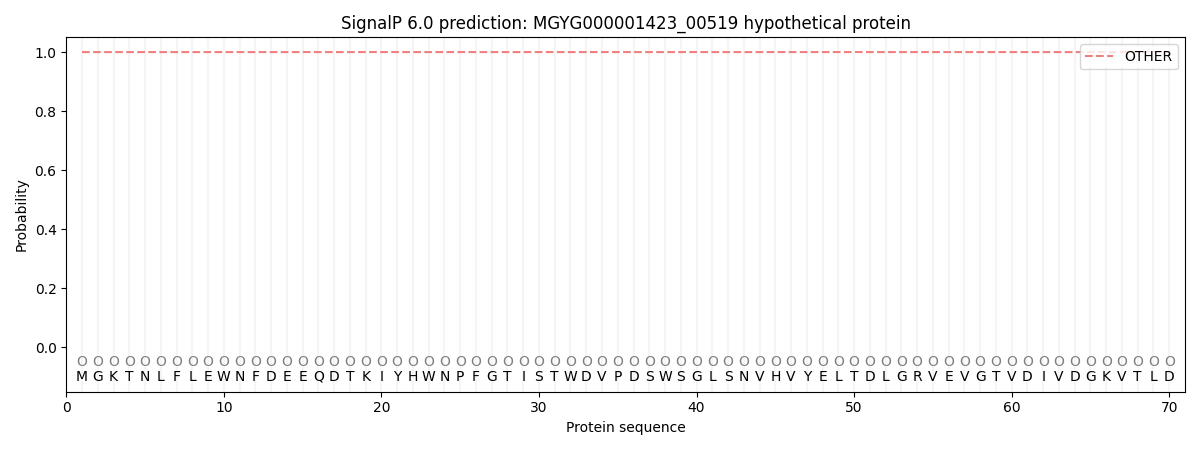
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000054 | 0.000004 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |