You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001432_00991
You are here: Home > Sequence: MGYG000001432_00991
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Helicobacter_C cinaedi | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Campylobacterota; Campylobacteria; Campylobacterales; Helicobacteraceae; Helicobacter_C; Helicobacter_C cinaedi | |||||||||||
| CAZyme ID | MGYG000001432_00991 | |||||||||||
| CAZy Family | GT9 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 959439; End: 960572 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GT9 | 57 | 348 | 3.7e-56 | 0.9511111111111111 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd03789 | GT9_LPS_heptosyltransferase | 8.33e-62 | 1 | 359 | 15 | 272 | lipopolysaccharide heptosyltransferase and similar proteins. Lipopolysaccharide heptosyltransferase (2.4.99.B6) is involved in the biosynthesis of lipooligosaccharide (LOS). Lipopolysaccharide (LPS) is a major component of the outer membrane of gram-negative bacteria. LPS heptosyltransferase transfers heptose molecules from ADP-heptose to 3-deoxy-D-manno-octulosonic acid (KDO), a part of the inner core component of LPS. This family also contains lipopolysaccharide 1,2-N-acetylglucosaminetransferase EC 2.4.1.56 and belongs to the GT-B structural superfamily of glycoslytransferases, which have characteristic N- and C-terminal domains each containing a typical Rossmann fold. The two domains have high structural homology despite minimal sequence homology. The large cleft that separates the two domains includes the catalytic center and permits a high degree of flexibility. |
| COG0859 | RfaF | 1.92e-54 | 1 | 370 | 17 | 334 | ADP-heptose:LPS heptosyltransferase [Cell wall/membrane/envelope biogenesis]. |
| TIGR02195 | heptsyl_trn_II | 2.32e-48 | 1 | 367 | 15 | 334 | lipopolysaccharide heptosyltransferase II. This family consists of examples of ADP-heptose:LPS heptosyltransferase II, an enzyme of LPS inner core region biosynthesis. LPS, composed of lipid A, a core region, and O antigen, is found in the outer membrane of Gram-negative bacteria. [Cell envelope, Biosynthesis and degradation of surface polysaccharides and lipopolysaccharides] |
| pfam01075 | Glyco_transf_9 | 3.12e-40 | 65 | 346 | 14 | 245 | Glycosyltransferase family 9 (heptosyltransferase). Members of this family belong to glycosyltransferase family 9. Lipopolysaccharide is a major component of the outer leaflet of the outer membrane in Gram-negative bacteria. It is composed of three domains; lipid A, Core oligosaccharide and the O-antigen. All of these enzymes transfer heptose to the lipopolysaccharide core. |
| PRK10916 | PRK10916 | 1.44e-29 | 63 | 315 | 80 | 297 | ADP-heptose--LPS heptosyltransferase RfaF. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| BAM32232.1 | 3.24e-272 | 1 | 377 | 1 | 377 |
| QOQ91694.1 | 1.26e-271 | 1 | 377 | 37 | 413 |
| BAM12215.1 | 1.93e-264 | 1 | 377 | 1 | 377 |
| QOQ95893.1 | 2.13e-263 | 1 | 377 | 37 | 413 |
| BBB19867.1 | 4.30e-263 | 1 | 377 | 37 | 413 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1PSW_A | 8.10e-21 | 63 | 359 | 80 | 336 | Structureof E. coli ADP-heptose lps heptosyltransferase II [Escherichia coli] |
| 3TOV_A | 5.15e-07 | 199 | 316 | 187 | 298 | Thecrystal structure of the glycosyl transferase family 9 from Veillonella parvula DSM 2008 [Veillonella parvula DSM 2008],3TOV_B The crystal structure of the glycosyl transferase family 9 from Veillonella parvula DSM 2008 [Veillonella parvula DSM 2008] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P37692 | 7.68e-22 | 63 | 359 | 80 | 336 | ADP-heptose--LPS heptosyltransferase 2 OS=Escherichia coli (strain K12) OX=83333 GN=rfaF PE=1 SV=1 |
| P37421 | 5.02e-21 | 63 | 359 | 80 | 336 | ADP-heptose--LPS heptosyltransferase 2 OS=Salmonella typhimurium (strain LT2 / SGSC1412 / ATCC 700720) OX=99287 GN=rfaF PE=3 SV=1 |
| P45042 | 1.50e-19 | 197 | 315 | 180 | 297 | ADP-heptose--LPS heptosyltransferase 2 OS=Haemophilus influenzae (strain ATCC 51907 / DSM 11121 / KW20 / Rd) OX=71421 GN=rfaF PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
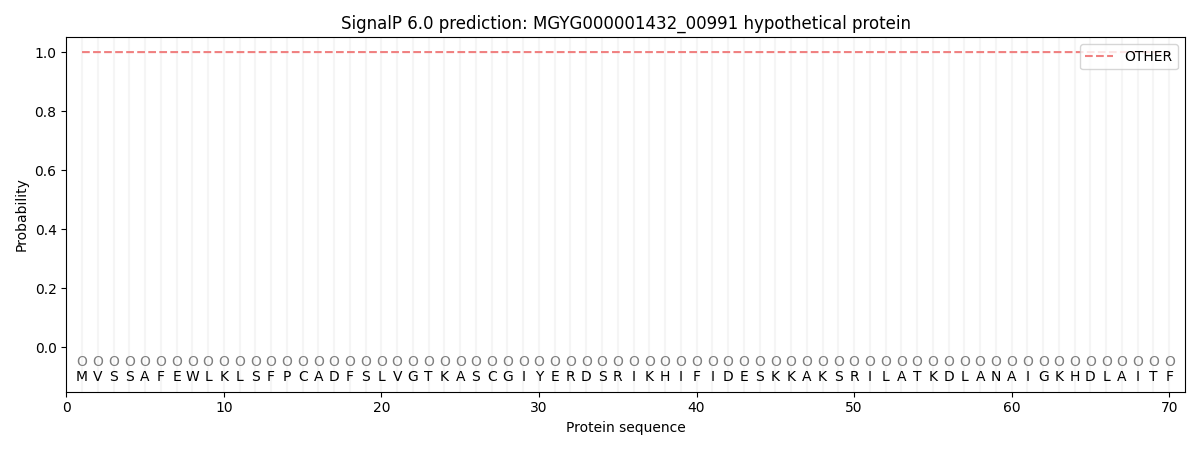
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000058 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
