You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001433_00952
You are here: Home > Sequence: MGYG000001433_00952
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Bacteroides salyersiae | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Bacteroidaceae; Bacteroides; Bacteroides salyersiae | |||||||||||
| CAZyme ID | MGYG000001433_00952 | |||||||||||
| CAZy Family | CE8 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 1164100; End: 1167888 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| CE8 | 779 | 1058 | 7.1e-70 | 0.8993055555555556 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| PLN02773 | PLN02773 | 4.20e-39 | 781 | 1051 | 9 | 263 | pectinesterase |
| pfam01095 | Pectinesterase | 1.12e-36 | 778 | 1052 | 1 | 260 | Pectinesterase. |
| PLN02432 | PLN02432 | 3.03e-35 | 781 | 1042 | 15 | 240 | putative pectinesterase |
| PLN02217 | PLN02217 | 1.43e-31 | 778 | 1106 | 251 | 551 | probable pectinesterase/pectinesterase inhibitor |
| COG4677 | PemB | 2.31e-30 | 756 | 1070 | 61 | 390 | Pectin methylesterase and related acyl-CoA thioesterases [Carbohydrate transport and metabolism, Lipid transport and metabolism]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QUT77822.1 | 0.0 | 1 | 1262 | 1 | 1262 |
| ADY51045.1 | 2.31e-192 | 345 | 1213 | 201 | 1070 |
| QCD39271.1 | 7.41e-192 | 324 | 1187 | 412 | 1247 |
| QCP72963.1 | 7.41e-192 | 324 | 1187 | 412 | 1247 |
| SCV07875.1 | 6.79e-190 | 672 | 1181 | 35 | 537 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1GQ8_A | 4.06e-22 | 778 | 1038 | 8 | 249 | Pectinmethylesterase from Carrot [Daucus carota] |
| 5C1E_A | 9.41e-22 | 781 | 1053 | 13 | 261 | CrystalStructure of the Pectin Methylesterase from Aspergillus niger in Penultimately Deglycosylated Form (N-acetylglucosamine Stub at Asn84) [Aspergillus niger ATCC 1015] |
| 5C1C_A | 1.27e-21 | 781 | 1053 | 13 | 261 | CrystalStructure of the Pectin Methylesterase from Aspergillus niger in Deglycosylated Form [Aspergillus niger ATCC 1015] |
| 4PMH_A | 3.30e-18 | 789 | 1016 | 48 | 292 | Thestructure of rice weevil pectin methyl esterase [Sitophilus oryzae] |
| 1XG2_A | 1.01e-17 | 776 | 1040 | 2 | 247 | ChainA, Pectinesterase 1 [Solanum lycopersicum] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q9LVQ0 | 2.60e-30 | 781 | 1051 | 9 | 263 | Pectinesterase 31 OS=Arabidopsis thaliana OX=3702 GN=PME31 PE=1 SV=1 |
| Q43043 | 1.24e-28 | 778 | 1104 | 58 | 359 | Pectinesterase OS=Petunia integrifolia OX=4103 GN=PPE1 PE=2 SV=1 |
| Q8GX86 | 5.62e-28 | 778 | 1106 | 255 | 555 | Probable pectinesterase/pectinesterase inhibitor 21 OS=Arabidopsis thaliana OX=3702 GN=PME21 PE=2 SV=2 |
| O80722 | 1.83e-27 | 780 | 1096 | 278 | 569 | Pectinesterase 4 OS=Arabidopsis thaliana OX=3702 GN=PME4 PE=2 SV=1 |
| Q9SMY6 | 2.14e-27 | 778 | 1093 | 296 | 585 | Putative pectinesterase/pectinesterase inhibitor 45 OS=Arabidopsis thaliana OX=3702 GN=PME45 PE=2 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
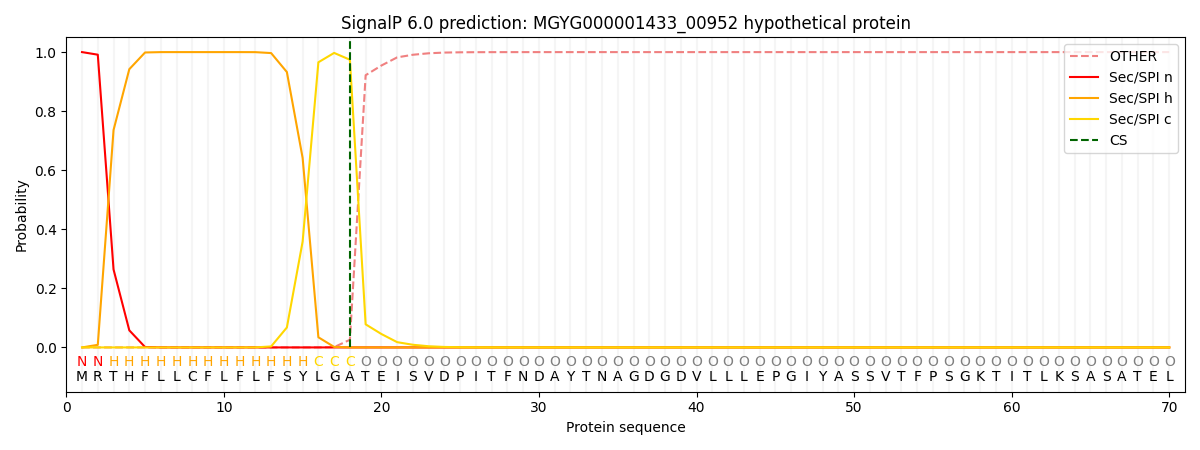
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000237 | 0.999151 | 0.000165 | 0.000152 | 0.000134 | 0.000131 |
