You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001436_00366
You are here: Home > Sequence: MGYG000001436_00366
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Paenibacillus_F sp000411255 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes; Bacilli; Paenibacillales; Paenibacillaceae; Paenibacillus_F; Paenibacillus_F sp000411255 | |||||||||||
| CAZyme ID | MGYG000001436_00366 | |||||||||||
| CAZy Family | PL6 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 469011; End: 472730 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| PL6 | 395 | 779 | 1.5e-128 | 0.9971910112359551 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd14251 | PL-6 | 7.66e-130 | 395 | 781 | 1 | 369 | Polysaccharide Lyase Family 6. Polysaccharide Lyase Family 6 is a family of beta-helical polysaccharide lyases. Members include alginate lyase (EC 4.2.2.3) and chondroitinase B (EC 4.2.2.19). Chondroitinase B is an enzyme that only cleaves the beta-(1,4)-linkage of dermatan sulfate (DS), leading to 4,5-unsaturated dermatan sulfate disaccharides as the product. DS is a highly sulfated, unbranched polysaccharide belonging to a family of glycosaminoglycans (GAGs) composed of alternating hexosamine (gluco- or galactosamine) and uronic acid (D-glucuronic or L-iduronic acid) moieties. DS contains alternating 1,4-beta-D-galactosamine (GalNac) and 1,3-alpha-L-iduronic acid units. The related chondroitin sulfate (CS) contains alternating GalNac and 1,3-beta-D-glucuronic acid units. Alginate lyases (known as either mannuronate (EC 4.2.2.3) or guluronate lyases (EC 4.2.2.11) catalyze the degradation of alginate, a copolymer of alpha-L-guluronate and its C5 epimer beta-D-mannuronate. |
| pfam14592 | Chondroitinas_B | 2.03e-49 | 395 | 778 | 1 | 380 | Chondroitinase B. This family includes chondroitinases. These enzymes cleave the glycosaminoglycan dermatan sulfate. |
| NF033679 | DNRLRE_dom | 1.54e-13 | 1078 | 1235 | 2 | 164 | DNRLRE domain. The DNRLRE domain, with a length of about 160 amino acids, appears typically in large, repetitive surface proteins of bacteria and archaea, sometimes repeated several times. It occurs, notably, three times in the C-terminal region of the enzyme disaggregatase from the archaeal species Methanosarcina mazei, each time with the motif DNRLRE, for which the domain is named. Archaeal proteins within this family are described particularly well by the currently more narrowly defined Pfam model, PF06848. Note that the catalytic region of disaggregatase, in the N-terminal portion of the protein, is modeled by a different HMM, PF08480. |
| cd00063 | FN3 | 3.09e-12 | 300 | 386 | 2 | 92 | Fibronectin type 3 domain; One of three types of internal repeats found in the plasma protein fibronectin. Its tenth fibronectin type III repeat contains an RGD cell recognition sequence in a flexible loop between 2 strands. Approximately 2% of all animal proteins contain the FN3 repeat; including extracellular and intracellular proteins, membrane spanning cytokine receptors, growth hormone receptors, tyrosine phosphatase receptors, and adhesion molecules. FN3-like domains are also found in bacterial glycosyl hydrolases. |
| cd00063 | FN3 | 1.71e-08 | 897 | 979 | 4 | 93 | Fibronectin type 3 domain; One of three types of internal repeats found in the plasma protein fibronectin. Its tenth fibronectin type III repeat contains an RGD cell recognition sequence in a flexible loop between 2 strands. Approximately 2% of all animal proteins contain the FN3 repeat; including extracellular and intracellular proteins, membrane spanning cytokine receptors, growth hormone receptors, tyrosine phosphatase receptors, and adhesion molecules. FN3-like domains are also found in bacterial glycosyl hydrolases. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QAV19742.1 | 0.0 | 1 | 1239 | 1 | 1239 |
| AIQ11428.1 | 9.28e-105 | 357 | 881 | 5 | 476 |
| QHY94407.1 | 6.25e-93 | 395 | 880 | 55 | 472 |
| QFI43533.1 | 5.42e-90 | 395 | 880 | 55 | 472 |
| CAB61820.1 | 5.42e-90 | 395 | 880 | 55 | 472 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 7O77_A | 1.05e-36 | 400 | 789 | 5 | 381 | ChainA, Poly(Beta-D-mannuronate) lyase [Pseudoalteromonas atlantica T6c] |
| 7O7T_A | 1.05e-36 | 400 | 789 | 5 | 381 | ChainA, Poly(Beta-D-mannuronate) lyase [Pseudoalteromonas atlantica T6c] |
| 5GKD_A | 7.07e-33 | 405 | 778 | 15 | 371 | Structureof PL6 family alginate lyase AlyGC [Paraglaciecola chathamensis],5GKD_B Structure of PL6 family alginate lyase AlyGC [Paraglaciecola chathamensis],5GKD_C Structure of PL6 family alginate lyase AlyGC [Paraglaciecola chathamensis],5GKD_D Structure of PL6 family alginate lyase AlyGC [Paraglaciecola chathamensis] |
| 5GKQ_A | 3.82e-32 | 405 | 778 | 15 | 371 | Structureof PL6 family alginate lyase AlyGC mutant-R241A [Paraglaciecola chathamensis S18K6],5GKQ_B Structure of PL6 family alginate lyase AlyGC mutant-R241A [Paraglaciecola chathamensis S18K6] |
| 6QPS_A | 4.11e-28 | 410 | 776 | 45 | 402 | Structuralcharacterization of a mannuronic acid specific polysaccharide family 6 lyase enzyme from human gut microbiota [Bacteroides cellulosilyticus],6QPS_B Structural characterization of a mannuronic acid specific polysaccharide family 6 lyase enzyme from human gut microbiota [Bacteroides cellulosilyticus] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q06365 | 4.68e-31 | 409 | 789 | 1 | 353 | Alginate lyase OS=Pseudomonas sp. (strain OS-ALG-9) OX=86038 GN=aly PE=3 SV=1 |
| Q46079 | 1.90e-13 | 410 | 860 | 43 | 467 | Chondroitinase-B OS=Pedobacter heparinus (strain ATCC 13125 / DSM 2366 / CIP 104194 / JCM 7457 / NBRC 12017 / NCIMB 9290 / NRRL B-14731 / HIM 762-3) OX=485917 GN=cslB PE=1 SV=2 |
| P20533 | 4.82e-08 | 819 | 984 | 396 | 554 | Chitinase A1 OS=Niallia circulans OX=1397 GN=chiA1 PE=1 SV=1 |
| P36909 | 3.05e-07 | 890 | 997 | 140 | 244 | Chitinase C OS=Streptomyces lividans OX=1916 GN=chiC PE=2 SV=1 |
| P11220 | 1.19e-06 | 850 | 979 | 101 | 226 | Chitinase 63 OS=Streptomyces plicatus OX=1922 GN=chtA PE=1 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as SP
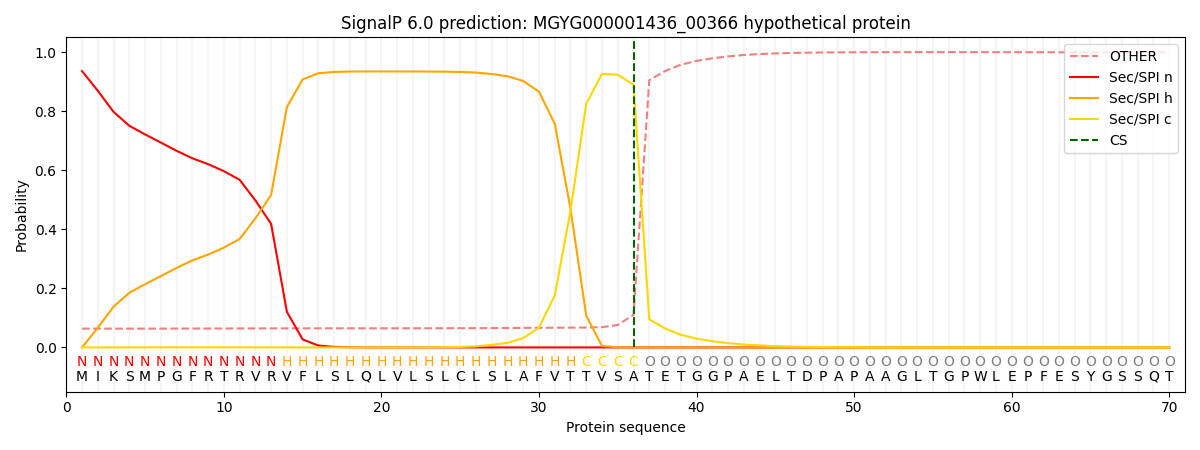
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.069300 | 0.928092 | 0.001149 | 0.000792 | 0.000342 | 0.000302 |

