You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001436_05516
You are here: Home > Sequence: MGYG000001436_05516
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Paenibacillus_F sp000411255 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes; Bacilli; Paenibacillales; Paenibacillaceae; Paenibacillus_F; Paenibacillus_F sp000411255 | |||||||||||
| CAZyme ID | MGYG000001436_05516 | |||||||||||
| CAZy Family | GT2 | |||||||||||
| CAZyme Description | D-alanine--poly(phosphoribitol) ligase subunit 1 | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 2814574; End: 2824830 Strand: - | |||||||||||
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd00833 | PKS | 0.0 | 31 | 453 | 2 | 421 | polyketide synthases (PKSs) polymerize simple fatty acids into a large variety of different products, called polyketides, by successive decarboxylating Claisen condensations. PKSs can be divided into 2 groups, modular type I PKSs consisting of one or more large multifunctional proteins and iterative type II PKSs, complexes of several monofunctional subunits. |
| PRK05691 | PRK05691 | 0.0 | 1298 | 3402 | 37 | 2154 | peptide synthase; Validated |
| PRK12316 | PRK12316 | 0.0 | 811 | 3401 | 1534 | 4064 | peptide synthase; Provisional |
| PRK12316 | PRK12316 | 0.0 | 1872 | 3402 | 1547 | 3023 | peptide synthase; Provisional |
| PRK12467 | PRK12467 | 0.0 | 1865 | 3402 | 32 | 1540 | peptide synthase; Provisional |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QND46664.1 | 1.03e-311 | 686 | 2954 | 442 | 2660 |
| BAY30132.1 | 8.98e-251 | 1 | 1867 | 1180 | 3301 |
| BAZ00088.1 | 2.89e-250 | 1 | 1867 | 1178 | 3299 |
| BAZ75991.1 | 2.89e-250 | 1 | 1867 | 1178 | 3299 |
| BAY90071.1 | 3.53e-250 | 24 | 1867 | 1205 | 3290 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6MFZ_A | 6.27e-290 | 1262 | 2965 | 186 | 1806 | Crystalstructure of dimodular LgrA in a condensation state [Brevibacillus parabrevis],6MFZ_B Crystal structure of dimodular LgrA in a condensation state [Brevibacillus parabrevis] |
| 6MFY_A | 8.29e-273 | 1262 | 2883 | 186 | 1725 | Crystalstructure of a 5-domain construct of LgrA in the substrate donation state [Brevibacillus parabrevis],6MG0_A Crystal structure of a 5-domain construct of LgrA in the thiolation state [Brevibacillus parabrevis],6MG0_B Crystal structure of a 5-domain construct of LgrA in the thiolation state [Brevibacillus parabrevis] |
| 5U89_A | 1.38e-176 | 2376 | 3400 | 26 | 1049 | Crystalstructure of a cross-module fragment from the dimodular NRPS DhbF [Geobacillus sp. Y4.1MC1] |
| 6MFW_A | 8.72e-174 | 2376 | 3382 | 207 | 1170 | Crystalstructure of a 4-domain construct of LgrA in the substrate donation state [Brevibacillus parabrevis] |
| 6MFX_A | 1.33e-172 | 2376 | 3382 | 207 | 1170 | Crystalstructure of a 4-domain construct of a mutant of LgrA in the substrate donation state [Brevibacillus parabrevis] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P39847 | 0.0 | 834 | 3289 | 8 | 2410 | Plipastatin synthase subunit C OS=Bacillus subtilis (strain 168) OX=224308 GN=ppsC PE=1 SV=2 |
| Q04747 | 0.0 | 830 | 3412 | 3 | 2516 | Surfactin synthase subunit 2 OS=Bacillus subtilis (strain 168) OX=224308 GN=srfAB PE=1 SV=3 |
| P94459 | 0.0 | 825 | 3254 | 1048 | 3402 | Plipastatin synthase subunit D OS=Bacillus subtilis (strain 168) OX=224308 GN=ppsD PE=1 SV=2 |
| Q70LM4 | 0.0 | 688 | 3400 | 968 | 4063 | Linear gramicidin synthase subunit D OS=Brevibacillus parabrevis OX=54914 GN=lgrD PE=1 SV=1 |
| P39846 | 0.0 | 834 | 3254 | 8 | 2366 | Plipastatin synthase subunit B OS=Bacillus subtilis (strain 168) OX=224308 GN=ppsB PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
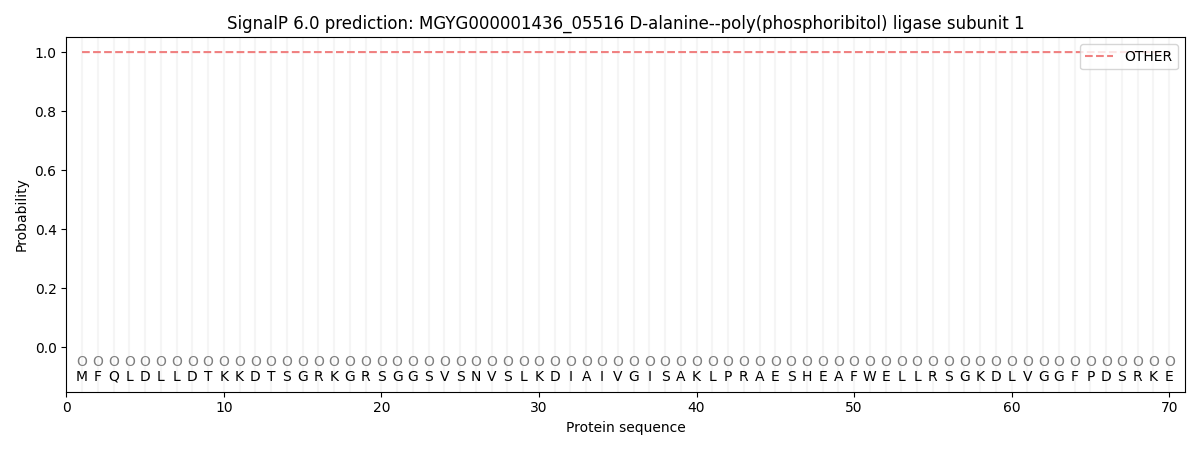
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000076 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
