You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001443_04741
You are here: Home > Sequence: MGYG000001443_04741
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Streptomyces albus | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Actinobacteriota; Actinomycetia; Streptomycetales; Streptomycetaceae; Streptomyces; Streptomyces albus | |||||||||||
| CAZyme ID | MGYG000001443_04741 | |||||||||||
| CAZy Family | GH46 | |||||||||||
| CAZyme Description | Chitosanase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 2626822; End: 2627667 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH46 | 48 | 265 | 4.5e-90 | 0.990990990990991 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam01374 | Glyco_hydro_46 | 2.24e-142 | 60 | 269 | 1 | 210 | Glycosyl hydrolase family 46. This family are chitosanase enzymes. |
| cd00978 | chitosanase_GH46 | 7.88e-106 | 52 | 269 | 1 | 222 | chitosanase belonging to the glycosyl hydrolase 46 family. This family is composed of the chitosanase enzymes which hydrolyzes chitosan, a biopolymer of beta (1,4)-linked-D-glucosamine (GlcN) residues produced by partial or full deacetylation of chitin. Chitosanases play a role in defense against pathogens such as fungi and are found in microorganisms, fungi, viruses, and plants. Microbial chitosanases can be divided into 3 subclasses based on the specificity of the cleavage positions for partial acetylated chitosan. Subclass I chitosanases such as N174 can split GlcN-GlcN and GlcNAc-GlcN linkages, whereas subclass II chitosanases such as Bacillus sp. no. 7-M can cleave only GlcN-GlcN linkages. Subclass III chitosanases such as MH-K1 chitosanase are the most versatile and can split both GlcN-GlcN and GlcN-GlcNAc linkages. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QID39641.1 | 3.46e-205 | 1 | 281 | 1 | 281 |
| AZP22132.1 | 1.96e-154 | 10 | 280 | 4 | 271 |
| CCK25255.1 | 2.41e-154 | 15 | 280 | 6 | 267 |
| ALV31577.1 | 2.71e-154 | 15 | 280 | 14 | 280 |
| QUC63506.1 | 5.62e-154 | 15 | 280 | 9 | 271 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 4ILY_A | 1.11e-140 | 30 | 280 | 3 | 249 | Abundantlysecreted chitosanase from Streptomyces sp. SirexAA-E [Streptomyces sp. SirexAA-E],4ILY_B Abundantly secreted chitosanase from Streptomyces sp. SirexAA-E [Streptomyces sp. SirexAA-E] |
| 1CHK_A | 1.54e-128 | 46 | 281 | 2 | 237 | StreptomycesN174 Chitosanase Ph5.5 298k [Streptomyces sp. N174],1CHK_B Streptomyces N174 Chitosanase Ph5.5 298k [Streptomyces sp. N174] |
| 4OLT_A | 1.00e-100 | 49 | 281 | 16 | 248 | Chitosanasecomplex structure [Pseudomonas sp. LL2(2010)],4OLT_B Chitosanase complex structure [Pseudomonas sp. LL2(2010)] |
| 4QWP_A | 1.65e-99 | 49 | 281 | 16 | 248 | co-crystalstructure of chitosanase OU01 with substrate [Pseudomonas sp. A-01],4QWP_B co-crystal structure of chitosanase OU01 with substrate [Pseudomonas sp. A-01] |
| 7C6C_A | 1.01e-38 | 47 | 269 | 1 | 229 | ChainA, Chitosanase [Bacillus subtilis subsp. subtilis str. 168] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P33665 | 3.12e-130 | 14 | 281 | 4 | 277 | Chitosanase OS=Streptomyces sp. (strain N174) OX=69019 GN=csn PE=1 SV=1 |
| P48846 | 2.91e-121 | 13 | 281 | 3 | 277 | Chitosanase OS=Nocardioides sp. (strain N106) OX=47919 GN=csn PE=3 SV=1 |
| O07921 | 1.34e-37 | 47 | 269 | 36 | 264 | Chitosanase OS=Bacillus subtilis (strain 168) OX=224308 GN=csn PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as TAT
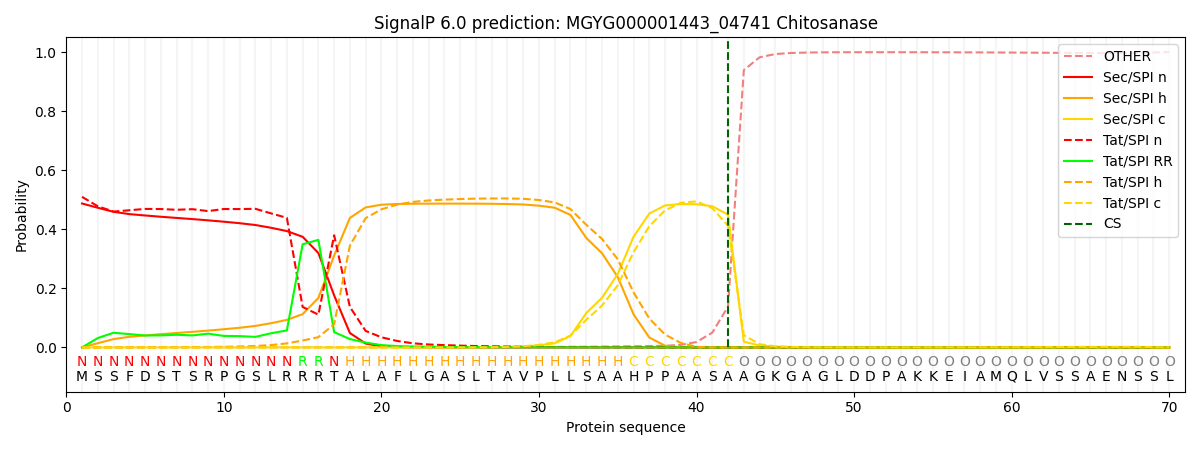
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.001416 | 0.482543 | 0.000560 | 0.512710 | 0.002583 | 0.000171 |
