You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001468_02601
You are here: Home > Sequence: MGYG000001468_02601
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
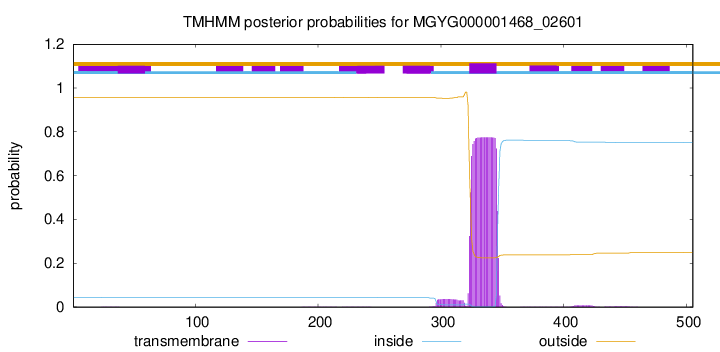
TMHMM annotations
Basic Information help
| Species | Blastococcus massiliensis | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Actinobacteriota; Actinomycetia; Mycobacteriales; Geodermatophilaceae; Blastococcus; Blastococcus massiliensis | |||||||||||
| CAZyme ID | MGYG000001468_02601 | |||||||||||
| CAZy Family | GT2 | |||||||||||
| CAZyme Description | Putative mycofactocin biosynthesis glycosyltransferase MftF | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 895783; End: 897300 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GT2 | 88 | 202 | 1e-20 | 0.7235294117647059 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| TIGR03965 | mycofact_glyco | 1.76e-104 | 13 | 464 | 1 | 463 | mycofactocin system glycosyltransferase. Members of this protein family are putative glycosyltransferases, members of pfam00535 (glycosyl transferase family 2). Members appear mostly in the Actinobacteria, where they appear to be part of a system for converting a precursor peptide (TIGR03969) into a novel redox carrier designated mycofactocin. A radical SAM enzyme, TIGR03962, is a proposed to be a key maturase for mycofactocin. |
| cd04186 | GT_2_like_c | 3.71e-16 | 118 | 284 | 29 | 166 | Subfamily of Glycosyltransferase Family GT2 of unknown function. GT-2 includes diverse families of glycosyltransferases with a common GT-A type structural fold, which has two tightly associated beta/alpha/beta domains that tend to form a continuous central sheet of at least eight beta-strands. These are enzymes that catalyze the transfer of sugar moieties from activated donor molecules to specific acceptor molecules, forming glycosidic bonds. Glycosyltransferases have been classified into more than 90 distinct sequence based families. |
| COG1216 | GT2 | 2.51e-13 | 118 | 298 | 35 | 242 | Glycosyltransferase, GT2 family [Carbohydrate transport and metabolism]. |
| COG1215 | BcsA | 1.00e-11 | 119 | 295 | 88 | 277 | Glycosyltransferase, catalytic subunit of cellulose synthase and poly-beta-1,6-N-acetylglucosamine synthase [Cell motility]. |
| pfam00535 | Glycos_transf_2 | 4.12e-11 | 118 | 243 | 30 | 164 | Glycosyl transferase family 2. Diverse family, transferring sugar from UDP-glucose, UDP-N-acetyl- galactosamine, GDP-mannose or CDP-abequose, to a range of substrates including cellulose, dolichol phosphate and teichoic acids. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| ADB77459.1 | 1.03e-201 | 7 | 476 | 15 | 487 |
| QNG36663.1 | 6.38e-196 | 7 | 469 | 11 | 475 |
| CCH90751.1 | 1.23e-176 | 7 | 473 | 11 | 479 |
| CCG05593.1 | 1.28e-158 | 7 | 386 | 10 | 387 |
| QDQ96943.1 | 7.79e-133 | 7 | 465 | 5 | 484 |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P46370 | 1.15e-117 | 7 | 463 | 54 | 509 | Uncharacterized 55.3 kDa protein in thcA 5'region OS=Rhodococcus erythropolis OX=1833 PE=4 SV=1 |
| P9WMX0 | 2.76e-113 | 4 | 463 | 3 | 466 | Pre-mycofactocin glycosyltransferase OS=Mycobacterium tuberculosis (strain CDC 1551 / Oshkosh) OX=83331 GN=mftF PE=3 SV=1 |
| P9WMX1 | 2.76e-113 | 4 | 463 | 3 | 466 | Pre-mycofactocin glycosyltransferase OS=Mycobacterium tuberculosis (strain ATCC 25618 / H37Rv) OX=83332 GN=mftF PE=1 SV=1 |
| A0QSC1 | 1.24e-111 | 7 | 463 | 6 | 466 | Pre-mycofactocin glycosyltransferase OS=Mycolicibacterium smegmatis (strain ATCC 700084 / mc(2)155) OX=246196 GN=mftF PE=3 SV=1 |
| P0A598 | 8.65e-09 | 87 | 272 | 6 | 182 | Uncharacterized glycosyltransferase Mb0553 OS=Mycobacterium bovis (strain ATCC BAA-935 / AF2122/97) OX=233413 GN=BQ2027_MB0553 PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
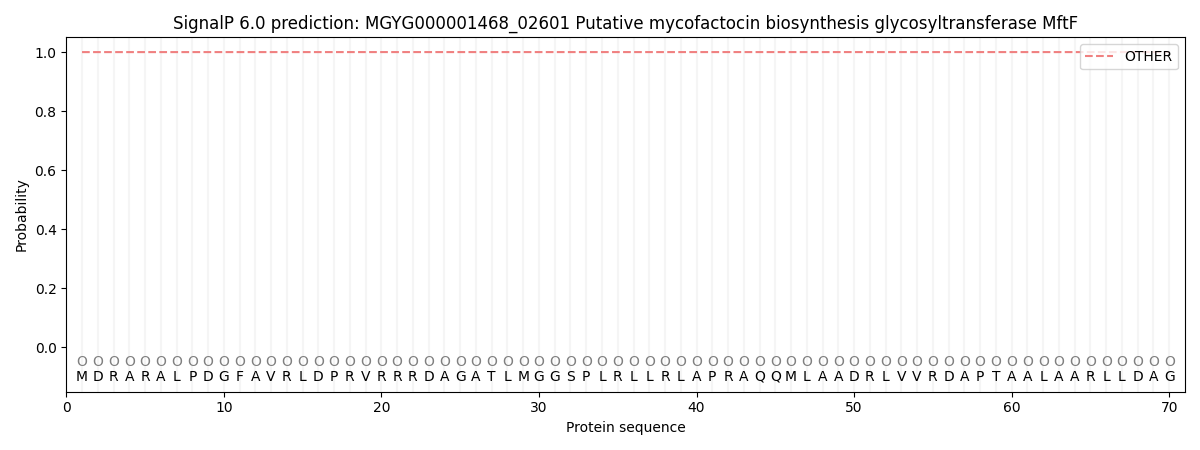
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000059 | 0.000001 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |