You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001468_03283
You are here: Home > Sequence: MGYG000001468_03283
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Blastococcus massiliensis | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Actinobacteriota; Actinomycetia; Mycobacteriales; Geodermatophilaceae; Blastococcus; Blastococcus massiliensis | |||||||||||
| CAZyme ID | MGYG000001468_03283 | |||||||||||
| CAZy Family | GH3 | |||||||||||
| CAZyme Description | Beta-hexosaminidase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 430126; End: 431331 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH3 | 131 | 353 | 6.6e-33 | 0.9351851851851852 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG1472 | BglX | 5.96e-34 | 72 | 372 | 4 | 293 | Periplasmic beta-glucosidase and related glycosidases [Carbohydrate transport and metabolism]. |
| pfam00933 | Glyco_hydro_3 | 1.26e-20 | 72 | 361 | 3 | 289 | Glycosyl hydrolase family 3 N terminal domain. |
| PRK05337 | PRK05337 | 3.10e-20 | 99 | 355 | 26 | 278 | beta-hexosaminidase; Provisional |
| PRK12438 | PRK12438 | 0.002 | 21 | 94 | 905 | 976 | hypothetical protein; Provisional |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QXG76235.1 | 2.02e-116 | 62 | 400 | 60 | 398 |
| ADB73175.1 | 3.07e-113 | 62 | 400 | 50 | 387 |
| QOK24004.1 | 9.05e-77 | 37 | 401 | 17 | 385 |
| QPP10539.1 | 5.00e-76 | 74 | 400 | 6 | 335 |
| QHC02406.1 | 7.06e-76 | 76 | 400 | 8 | 334 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5BU9_A | 4.07e-63 | 67 | 397 | 3 | 340 | Crystalstructure of Beta-N-acetylhexosaminidase from Beutenbergia cavernae DSM 12333 [Beutenbergia cavernae DSM 12333],5BU9_B Crystal structure of Beta-N-acetylhexosaminidase from Beutenbergia cavernae DSM 12333 [Beutenbergia cavernae DSM 12333] |
| 6GFV_A | 7.79e-24 | 69 | 367 | 16 | 310 | Mtuberculosis LpqI [Mycobacterium tuberculosis H37Rv],6GFV_B M tuberculosis LpqI [Mycobacterium tuberculosis H37Rv] |
| 3TEV_A | 4.37e-19 | 77 | 354 | 20 | 290 | Thecrystal structure of glycosyl hydrolase from Deinococcus radiodurans R1 [Deinococcus radiodurans R1],3TEV_B The crystal structure of glycosyl hydrolase from Deinococcus radiodurans R1 [Deinococcus radiodurans R1] |
| 4YYF_A | 5.03e-19 | 59 | 388 | 29 | 353 | ChainA, Beta-N-acetylhexosaminidase [Mycolicibacterium smegmatis MC2 155],4YYF_B Chain B, Beta-N-acetylhexosaminidase [Mycolicibacterium smegmatis MC2 155],4YYF_C Chain C, Beta-N-acetylhexosaminidase [Mycolicibacterium smegmatis MC2 155] |
| 6K5J_A | 1.25e-18 | 69 | 355 | 11 | 297 | Structureof a glycoside hydrolase family 3 beta-N-acetylglucosaminidase from Paenibacillus sp. str. FPU-7 [Paenibacillaceae] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| B0RX17 | 1.18e-25 | 78 | 372 | 1 | 293 | Beta-hexosaminidase OS=Xanthomonas campestris pv. campestris (strain B100) OX=509169 GN=nagZ PE=3 SV=1 |
| Q4USG7 | 1.18e-25 | 78 | 372 | 1 | 293 | Beta-hexosaminidase OS=Xanthomonas campestris pv. campestris (strain 8004) OX=314565 GN=nagZ PE=3 SV=1 |
| Q8PB42 | 1.18e-25 | 78 | 372 | 1 | 293 | Beta-hexosaminidase OS=Xanthomonas campestris pv. campestris (strain ATCC 33913 / DSM 3586 / NCPPB 528 / LMG 568 / P 25) OX=190485 GN=nagZ PE=3 SV=1 |
| B0U3L0 | 1.61e-24 | 97 | 383 | 22 | 304 | Beta-hexosaminidase OS=Xylella fastidiosa (strain M12) OX=405440 GN=nagZ PE=3 SV=1 |
| Q5H1Q0 | 1.06e-23 | 78 | 372 | 1 | 293 | Beta-hexosaminidase OS=Xanthomonas oryzae pv. oryzae (strain KACC10331 / KXO85) OX=291331 GN=nagZ PE=3 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as LIPO
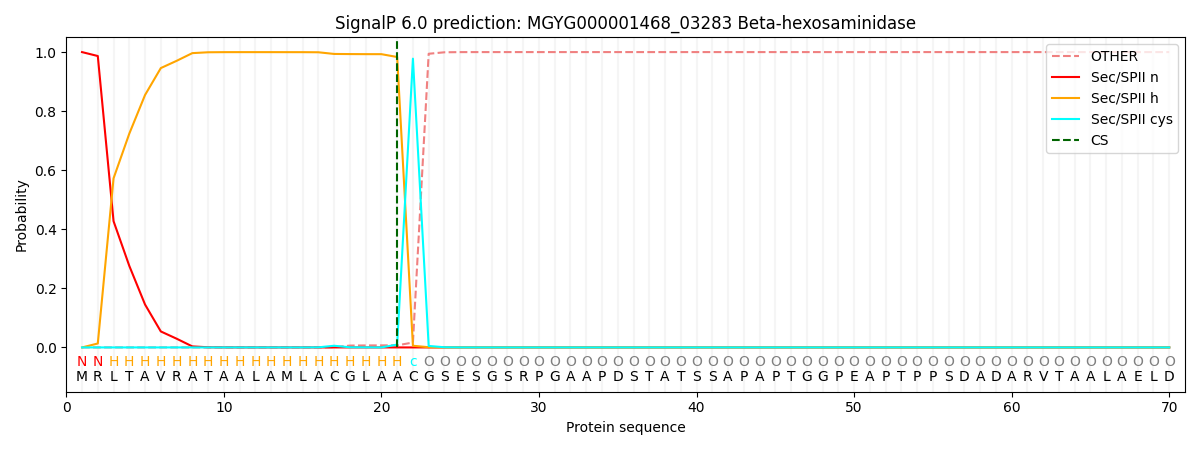
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000000 | 0.000007 | 1.000021 | 0.000000 | 0.000000 | 0.000000 |
