You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001476_01910
You are here: Home > Sequence: MGYG000001476_01910
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Caproiciproducens sp000752215 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Oscillospirales; Acutalibacteraceae; Caproiciproducens; Caproiciproducens sp000752215 | |||||||||||
| CAZyme ID | MGYG000001476_01910 | |||||||||||
| CAZy Family | GT83 | |||||||||||
| CAZyme Description | Undecaprenyl phosphate-alpha-4-amino-4-deoxy-L-arabinose arabinosyl transferase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 335404; End: 337542 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GT83 | 11 | 235 | 1.3e-37 | 0.40555555555555556 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG1807 | ArnT | 5.28e-43 | 16 | 574 | 9 | 446 | 4-amino-4-deoxy-L-arabinose transferase or related glycosyltransferase of PMT family [Cell wall/membrane/envelope biogenesis]. |
| pfam13231 | PMT_2 | 4.25e-34 | 70 | 224 | 1 | 157 | Dolichyl-phosphate-mannose-protein mannosyltransferase. This family contains members that are not captured by pfam02366. |
| COG4745 | COG4745 | 1.67e-07 | 85 | 242 | 79 | 253 | Predicted membrane-bound mannosyltransferase [General function prediction only]. |
| COG1928 | PMT1 | 3.79e-07 | 7 | 185 | 17 | 210 | Dolichyl-phosphate-mannose--protein O-mannosyl transferase [Posttranslational modification, protein turnover, chaperones]. |
| PRK13279 | arnT | 3.86e-06 | 71 | 232 | 62 | 227 | lipid IV(A) 4-amino-4-deoxy-L-arabinosyltransferase. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QHU89858.1 | 1.09e-60 | 36 | 685 | 294 | 889 |
| QHU90800.1 | 4.53e-57 | 22 | 661 | 280 | 859 |
| QJU08762.1 | 8.89e-55 | 22 | 685 | 279 | 895 |
| QUB37647.1 | 2.88e-54 | 17 | 686 | 210 | 781 |
| QHU92512.1 | 2.70e-52 | 36 | 661 | 292 | 859 |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P37483 | 1.33e-118 | 9 | 711 | 2 | 685 | Putative mannosyltransferase YycA OS=Bacillus subtilis (strain 168) OX=224308 GN=yycA PE=3 SV=2 |
| O34575 | 7.01e-106 | 9 | 694 | 4 | 673 | Putative mannosyltransferase YkcB OS=Bacillus subtilis (strain 168) OX=224308 GN=ykcB PE=3 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
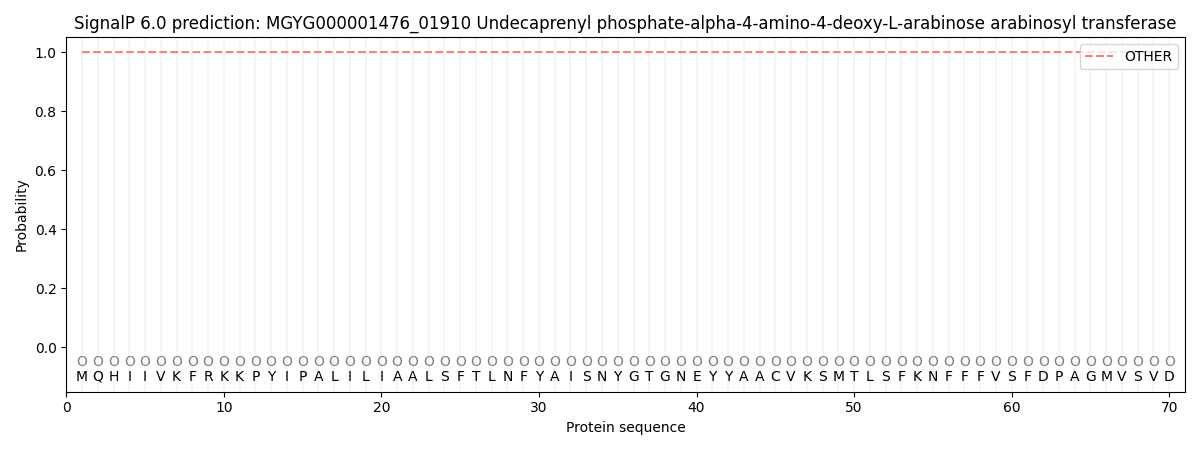
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.999966 | 0.000055 | 0.000001 | 0.000000 | 0.000000 | 0.000005 |
TMHMM Annotations download full data without filtering help
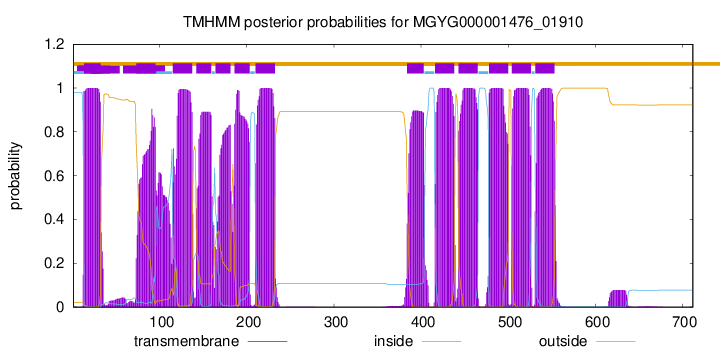
| start | end |
|---|---|
| 13 | 32 |
| 73 | 95 |
| 115 | 137 |
| 142 | 159 |
| 164 | 181 |
| 186 | 203 |
| 210 | 232 |
| 384 | 403 |
| 416 | 438 |
| 443 | 465 |
| 478 | 500 |
| 504 | 526 |
| 531 | 553 |
