You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001477_02740
You are here: Home > Sequence: MGYG000001477_02740
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Clostridium_L amazonitimonense | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes_A; Clostridia; Clostridiales; Clostridiaceae; Clostridium_L; Clostridium_L amazonitimonense | |||||||||||
| CAZyme ID | MGYG000001477_02740 | |||||||||||
| CAZy Family | CBM50 | |||||||||||
| CAZyme Description | Trifunctional nucleotide phosphoesterase protein YfkN | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 9549; End: 11129 Strand: - | |||||||||||
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG0737 | UshA | 8.52e-121 | 3 | 486 | 21 | 505 | 2',3'-cyclic-nucleotide 2'-phosphodiesterase/5'- or 3'-nucleotidase, 5'-nucleotidase family [Nucleotide transport and metabolism, Defense mechanisms]. |
| cd07410 | MPP_CpdB_N | 3.12e-110 | 9 | 288 | 1 | 278 | Escherichia coli CpdB and related proteins, N-terminal metallophosphatase domain. CpdB is a bacterial periplasmic protein with an N-terminal metallophosphatase domain and a C-terminal 3'-nucleotidase domain. This alignment model represents the N-terminal metallophosphatase domain, which has 2',3'-cyclic phosphodiesterase activity, hydrolyzing the 2',3'-cyclic phosphates of adenosine, guanosine, cytosine and uridine to yield nucleoside and phosphate. CpdB also hydrolyzes the chromogenic substrates p-nitrophenyl phosphate (PNPP), bis(PNPP) and p-nitrophenyl phosphorylcholine (NPPC). CpdB is thought to play a scavenging role during RNA hydrolysis by converting the non-transportable nucleotides produced by RNaseI to nucleosides which can easily enter a cell for use as a carbon source. This family also includes YfkN, a Bacillus subtilis nucleotide phosphoesterase with two copies of each of the metallophosphatase and 3'-nucleotidase domains. The N-terminal metallophosphatase domain belongs to a large superfamily of distantly related metallophosphatases (MPPs) that includes: Mre11/SbcD-like exonucleases, Dbr1-like RNA lariat debranching enzymes, YfcE-like phosphodiesterases, purple acid phosphatases (PAPs), YbbF-like UDP-2,3-diacylglucosamine hydrolases, and acid sphingomyelinases (ASMases). MPPs are functionally diverse, but all share a conserved domain with an active site consisting of two metal ions (usually manganese, iron, or zinc) coordinated with octahedral geometry by a cage of histidine, aspartate, and asparagine residues. The conserved domain is a double beta-sheet sandwich with a di-metal active site made up of residues located at the C-terminal side of the sheets. This domain is thought to allow for productive metal coordination. |
| PRK09419 | PRK09419 | 5.05e-107 | 8 | 525 | 41 | 603 | multifunctional 2',3'-cyclic-nucleotide 2'-phosphodiesterase/3'-nucleotidase/5'-nucleotidase. |
| PRK09420 | cpdB | 2.37e-86 | 11 | 522 | 28 | 595 | bifunctional 2',3'-cyclic-nucleotide 2'-phosphodiesterase/3'-nucleotidase. |
| TIGR01390 | CycNucDiestase | 7.26e-86 | 8 | 522 | 2 | 572 | 2',3'-cyclic-nucleotide 2'-phosphodiesterase. 2',3'-cyclic-nucleotide 2'-phosphodiesterase is a bifunctional enzyme localized to the periplasm of Gram-negative bacteria. 2',3'-cyclic-nucleotide 2'-phosphodiesters are intermediates formed during the hydrolysis of RNA by the ribonuclease I, which is also found to the periplasm, and other enzymes of the RNAse T2 family. Bacteria are unable to transport 2',3'-cyclic-nucleotides into the cytoplasm. 2',3'-cyclic-nucleotide 2'-phosphodiesterase contains 2 active sites which catalyze the reactions that convert the 2',3'-cyclic-nucleotide into a 3'-nucleotide, which is then converted into nucleic acid and phosphate. Both final products can be transported into the cytoplasm. Thus, it has been suggested that 2',3'-cyclic-nucleotide 2'-phosphodiesterase has a 'scavenging' function. Experimental evidence indicates that 2',3'-cyclic-nucleotide 2'-phosphodiesterase enables Yersinia enterocolitica O:8 to grow on 2'3'-cAMP as a sole source of carbon and energy (). [Purines, pyrimidines, nucleosides, and nucleotides, Other] |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| APT74155.1 | 4.49e-176 | 9 | 525 | 21 | 532 |
| QTA38048.1 | 1.69e-172 | 8 | 525 | 23 | 539 |
| ANQ54130.1 | 9.94e-170 | 8 | 525 | 20 | 532 |
| APT72575.1 | 9.94e-170 | 8 | 525 | 20 | 532 |
| AGB41602.1 | 2.30e-100 | 9 | 524 | 37 | 541 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3QFK_A | 1.70e-94 | 9 | 515 | 20 | 516 | ChainA, Uncharacterized protein [Staphylococcus aureus subsp. aureus NCTC 8325] |
| 4Q7F_A | 1.70e-94 | 9 | 515 | 20 | 516 | ChainA, 5' nucleotidase family protein [Staphylococcus aureus subsp. aureus COL] |
| 3GVE_A | 3.24e-40 | 11 | 282 | 14 | 305 | Crystalstructure of calcineurin-like phosphoesterase YfkN from Bacillus subtilis [Bacillus subtilis subsp. subtilis str. 168],3GVE_B Crystal structure of calcineurin-like phosphoesterase YfkN from Bacillus subtilis [Bacillus subtilis subsp. subtilis str. 168] |
| 3IVD_A | 5.77e-37 | 9 | 487 | 7 | 459 | Putative5'-Nucleotidase (c4898) from Escherichia Coli in complex with Uridine [Escherichia coli O6],3IVD_B Putative 5'-Nucleotidase (c4898) from Escherichia Coli in complex with Uridine [Escherichia coli O6],3IVE_A Putative 5'-Nucleotidase (c4898) from Escherichia Coli in complex with Cytidine [Escherichia coli O6] |
| 5EQV_A | 3.74e-36 | 11 | 303 | 12 | 327 | 1.45Angstrom Crystal Structure of Bifunctional 2',3'-cyclic Nucleotide 2'-phosphodiesterase/3'-Nucleotidase Periplasmic Precursor Protein from Yersinia pestis with Phosphate bound to the Active site [Yersinia pestis CO92] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| O34313 | 1.30e-72 | 11 | 524 | 47 | 609 | Trifunctional nucleotide phosphoesterase protein YfkN OS=Bacillus subtilis (strain 168) OX=224308 GN=yfkN PE=1 SV=1 |
| P44764 | 8.34e-65 | 11 | 525 | 36 | 606 | 2',3'-cyclic-nucleotide 2'-phosphodiesterase/3'-nucleotidase OS=Haemophilus influenzae (strain ATCC 51907 / DSM 11121 / KW20 / Rd) OX=71421 GN=cpdB PE=3 SV=1 |
| P08331 | 9.76e-63 | 11 | 524 | 26 | 595 | 2',3'-cyclic-nucleotide 2'-phosphodiesterase/3'-nucleotidase OS=Escherichia coli (strain K12) OX=83333 GN=cpdB PE=1 SV=2 |
| P26265 | 2.62e-62 | 11 | 524 | 26 | 595 | 2',3'-cyclic-nucleotide 2'-phosphodiesterase/3'-nucleotidase OS=Salmonella typhimurium (strain LT2 / SGSC1412 / ATCC 700720) OX=99287 GN=cpdB PE=3 SV=2 |
| P53052 | 3.61e-56 | 11 | 524 | 31 | 600 | 2',3'-cyclic-nucleotide 2'-phosphodiesterase/3'-nucleotidase OS=Yersinia enterocolitica OX=630 GN=cpdB PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
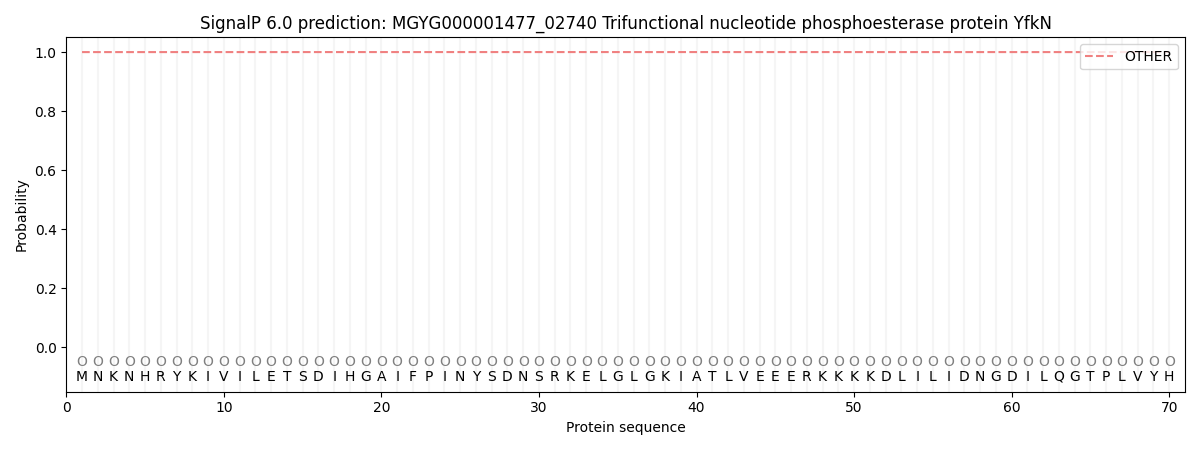
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000088 | 0.000001 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
