You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001483_00824
You are here: Home > Sequence: MGYG000001483_00824
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Arachnia massiliensis | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Actinobacteriota; Actinomycetia; Propionibacteriales; Propionibacteriaceae; Arachnia; Arachnia massiliensis | |||||||||||
| CAZyme ID | MGYG000001483_00824 | |||||||||||
| CAZy Family | CBM50 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 255969; End: 256673 Strand: + | |||||||||||
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| smart01043 | BTAD | 5.63e-06 | 85 | 214 | 2 | 137 | Bacterial transcriptional activator domain. Found in the DNRI/REDD/AFSR family of regulators. This region of AFSR along with the C terminal region is capable of independently directing actinorhodin production. This family contains TPR repeats. |
| COG4948 | RspA | 0.008 | 124 | 231 | 63 | 190 | L-alanine-DL-glutamate epimerase or related enzyme of enolase superfamily [Cell wall/membrane/envelope biogenesis, General function prediction only]. |
| pfam03704 | BTAD | 0.008 | 90 | 217 | 14 | 139 | Bacterial transcriptional activator domain. Found in the DNRI/REDD/AFSR family of regulators. This region of AFSR, along with the C terminal region, is capable of independently directing actinorhodin production. This family contains TPR repeats. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QTO37425.1 | 2.79e-120 | 1 | 227 | 665 | 891 |
| AQP50973.1 | 2.29e-97 | 1 | 226 | 671 | 896 |
| AQP45822.1 | 4.14e-94 | 1 | 223 | 664 | 885 |
| VEP40458.1 | 8.80e-91 | 1 | 222 | 652 | 873 |
| AQX15971.1 | 8.80e-91 | 1 | 222 | 652 | 873 |
Swiss-Prot Hits help
SignalP and Lipop Annotations help
This protein is predicted as OTHER
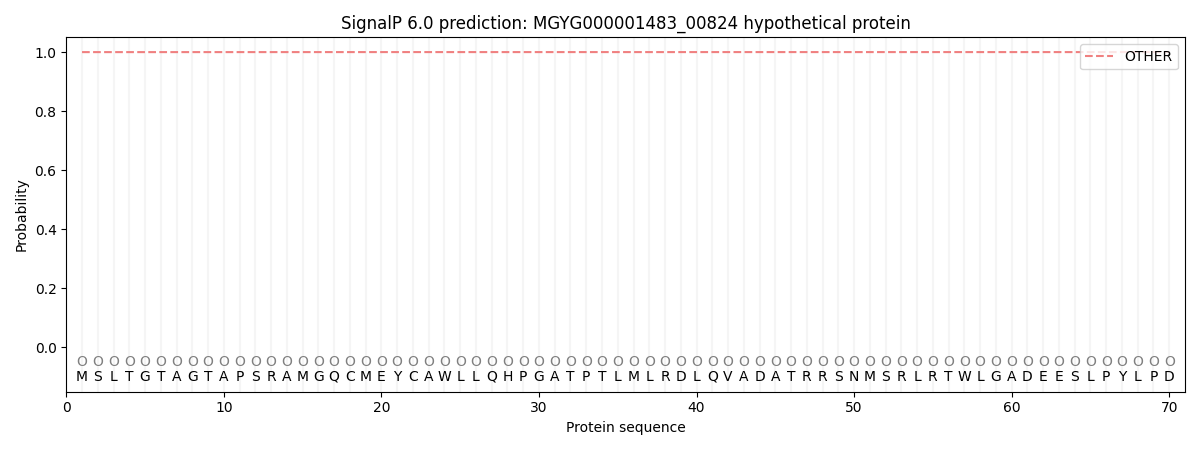
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000032 | 0.000001 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
