You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001484_00570
You are here: Home > Sequence: MGYG000001484_00570
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Nigerium massiliense | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Actinobacteriota; Actinomycetia; Propionibacteriales; Propionibacteriaceae; Nigerium; Nigerium massiliense | |||||||||||
| CAZyme ID | MGYG000001484_00570 | |||||||||||
| CAZy Family | GH28 | |||||||||||
| CAZyme Description | Endo-polygalacturonase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 297932; End: 299380 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH28 | 61 | 454 | 9.3e-73 | 0.9661538461538461 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG5434 | Pgu1 | 8.33e-51 | 42 | 377 | 88 | 402 | Polygalacturonase [Carbohydrate transport and metabolism]. |
| PLN02188 | PLN02188 | 5.99e-34 | 41 | 458 | 41 | 394 | polygalacturonase/glycoside hydrolase family protein |
| pfam00295 | Glyco_hydro_28 | 1.65e-25 | 214 | 455 | 85 | 316 | Glycosyl hydrolases family 28. Glycosyl hydrolase family 28 includes polygalacturonase EC:3.2.1.15 as well as rhamnogalacturonase A(RGase A), EC:3.2.1.-. These enzymes are important in cell wall metabolism. |
| PLN02218 | PLN02218 | 4.36e-23 | 42 | 383 | 73 | 355 | polygalacturonase ADPG |
| PLN03010 | PLN03010 | 2.49e-21 | 41 | 384 | 51 | 321 | polygalacturonase |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| APT58997.1 | 2.18e-113 | 43 | 476 | 36 | 457 |
| AWV20899.1 | 1.86e-111 | 41 | 476 | 63 | 478 |
| QET94657.1 | 2.05e-110 | 41 | 476 | 51 | 466 |
| ATR22288.1 | 2.05e-110 | 41 | 476 | 51 | 466 |
| AFW03263.1 | 5.79e-107 | 31 | 481 | 45 | 464 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1BHE_A | 2.39e-32 | 53 | 454 | 26 | 372 | ChainA, POLYGALACTURONASE [Pectobacterium carotovorum] |
| 3JUR_A | 2.29e-26 | 42 | 450 | 33 | 420 | Thecrystal structure of a hyperthermoactive Exopolygalacturonase from Thermotoga maritima [Thermotoga maritima],3JUR_B The crystal structure of a hyperthermoactive Exopolygalacturonase from Thermotoga maritima [Thermotoga maritima],3JUR_C The crystal structure of a hyperthermoactive Exopolygalacturonase from Thermotoga maritima [Thermotoga maritima],3JUR_D The crystal structure of a hyperthermoactive Exopolygalacturonase from Thermotoga maritima [Thermotoga maritima] |
| 2UVE_A | 6.61e-16 | 42 | 373 | 162 | 490 | Structureof Yersinia enterocolitica Family 28 Exopolygalacturonase [Yersinia enterocolitica],2UVE_B Structure of Yersinia enterocolitica Family 28 Exopolygalacturonase [Yersinia enterocolitica],2UVF_A Structure of Yersinia enterocolitica Family 28 Exopolygalacturonase in Complex with Digalaturonic Acid [Yersinia enterocolitica],2UVF_B Structure of Yersinia enterocolitica Family 28 Exopolygalacturonase in Complex with Digalaturonic Acid [Yersinia enterocolitica] |
| 6KVE_A | 1.85e-12 | 207 | 450 | 101 | 336 | ChainA, Endo-polygalacturonase [Evansstolkia leycettana] |
| 7E56_A | 4.18e-12 | 207 | 450 | 93 | 328 | ChainA, Endo-polygalacturonase [Evansstolkia leycettana] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P18192 | 4.14e-32 | 53 | 455 | 52 | 399 | Endo-polygalacturonase OS=Pectobacterium carotovorum subsp. carotovorum OX=555 GN=peh PE=3 SV=1 |
| P26509 | 2.02e-31 | 53 | 454 | 52 | 398 | Endo-polygalacturonase OS=Pectobacterium parmentieri OX=1905730 GN=pehA PE=1 SV=1 |
| A7PZL3 | 1.69e-30 | 42 | 467 | 68 | 453 | Probable polygalacturonase OS=Vitis vinifera OX=29760 GN=GSVIVT00026920001 PE=1 SV=1 |
| P35339 | 1.99e-23 | 41 | 446 | 45 | 386 | Exopolygalacturonase OS=Zea mays OX=4577 GN=PG2C PE=2 SV=1 |
| P58598 | 3.76e-23 | 46 | 383 | 68 | 401 | Polygalacturonase OS=Ralstonia solanacearum (strain GMI1000) OX=267608 GN=pglA PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
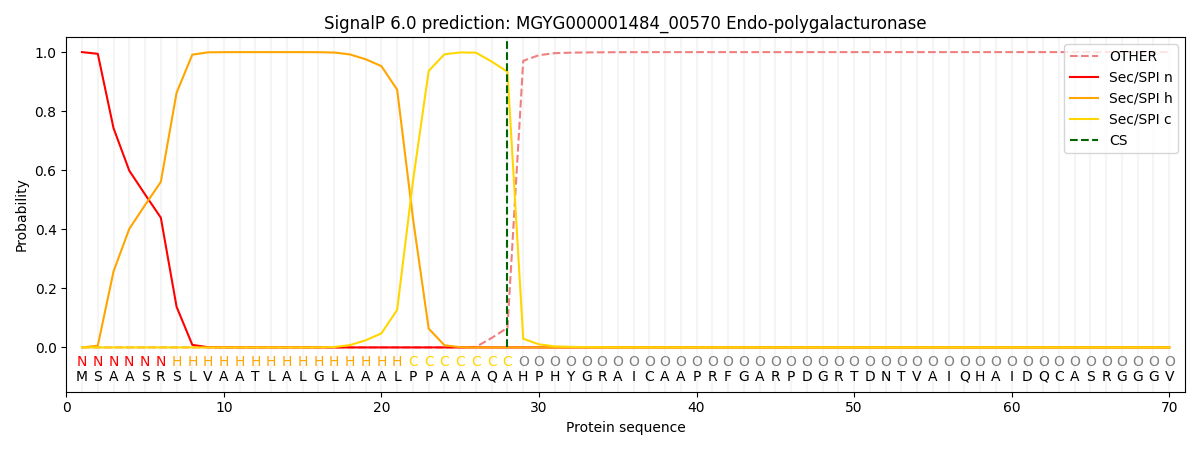
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000304 | 0.998959 | 0.000158 | 0.000236 | 0.000163 | 0.000149 |