You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001485_00422
Basic Information
help
| Species |
Pseudomonas_E massiliensis
|
| Lineage |
Bacteria; Proteobacteria; Gammaproteobacteria; Pseudomonadales; Pseudomonadaceae; Pseudomonas_E; Pseudomonas_E massiliensis
|
| CAZyme ID |
MGYG000001485_00422
|
| CAZy Family |
GT73 |
| CAZyme Description |
hypothetical protein
|
| CAZyme Property |
| Protein Length |
CGC |
Molecular Weight |
Isoelectric Point |
| 264 |
|
29570.89 |
9.8595 |
|
| Genome Property |
| Genome Assembly ID |
Genome Size |
Genome Type |
Country |
Continent |
| MGYG000001485 |
4723534 |
Isolate |
not provided |
not provided |
|
| Gene Location |
Start: 142834;
End: 143628
Strand: -
|
No EC number prediction in MGYG000001485_00422.
| Family |
Start |
End |
Evalue |
family coverage |
| GT73 |
18 |
246 |
9.1e-61 |
0.9755102040816327 |
| Cdd ID |
Domain |
E-Value |
qStart |
qEnd |
sStart |
sEnd |
Domain Description |
| PRK09822
|
PRK09822 |
6.03e-22 |
22 |
255 |
24 |
264 |
lipopolysaccharide core biosynthesis protein; Provisional |
| pfam01973
|
MAF_flag10 |
5.04e-06 |
21 |
182 |
27 |
166 |
Protein of unknown function DUF115. This family of archaebacterial proteins has no known function. |
| Hit ID |
E-Value |
Query Start |
Query End |
Hit Start |
Hit End |
Description |
|
P27241
|
1.24e-17 |
22 |
243 |
24 |
252 |
Lipopolysaccharide core biosynthesis protein RfaZ OS=Escherichia coli (strain K12) OX=83333 GN=rfaZ PE=4 SV=2 |
|
P26473
|
1.84e-16 |
23 |
255 |
25 |
264 |
Lipopolysaccharide core biosynthesis protein RfaZ OS=Salmonella typhimurium (strain LT2 / SGSC1412 / ATCC 700720) OX=99287 GN=rfaZ PE=4 SV=1 |
This protein is predicted as OTHER
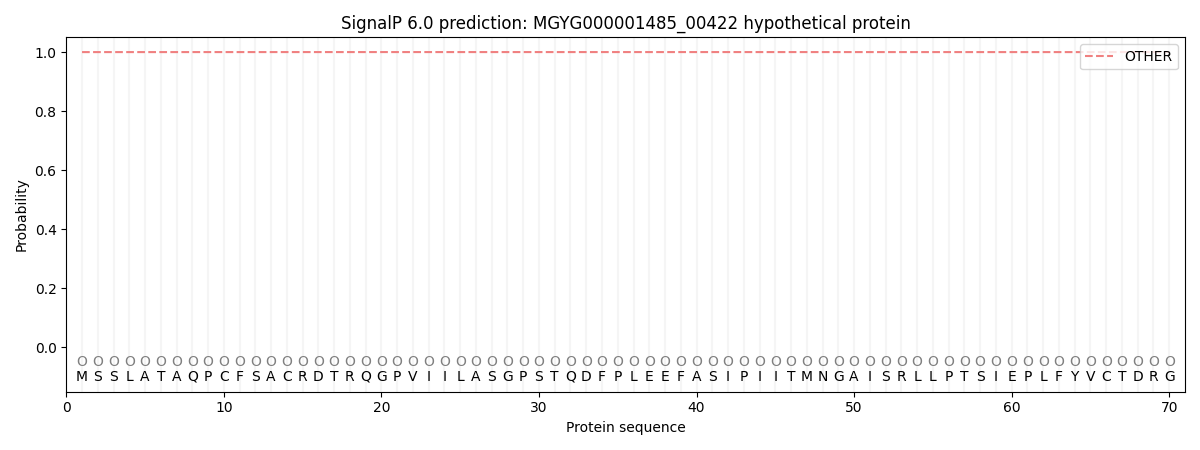
| Other |
SP_Sec_SPI |
LIPO_Sec_SPII |
TAT_Tat_SPI |
TATLIP_Sec_SPII |
PILIN_Sec_SPIII |
|
1.000061
|
0.000003
|
0.000000
|
0.000000
|
0.000000
|
0.000000
|
There is no transmembrane helices in MGYG000001485_00422.

