You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001507_03134
You are here: Home > Sequence: MGYG000001507_03134
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Paenibacillus ihuae | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes; Bacilli; Paenibacillales; Paenibacillaceae; Paenibacillus; Paenibacillus ihuae | |||||||||||
| CAZyme ID | MGYG000001507_03134 | |||||||||||
| CAZy Family | GH5 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 3283577; End: 3285043 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH5 | 42 | 287 | 5e-100 | 0.9841897233201581 |
| CBM59 | 343 | 486 | 1e-58 | 0.9862068965517241 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam00150 | Cellulase | 5.12e-42 | 41 | 285 | 1 | 272 | Cellulase (glycosyl hydrolase family 5). |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| BAA25878.1 | 7.44e-286 | 2 | 488 | 7 | 516 |
| AIQ62043.1 | 4.10e-280 | 10 | 488 | 3 | 485 |
| QUL52210.1 | 1.74e-271 | 1 | 488 | 5 | 520 |
| AAS48170.1 | 1.53e-218 | 6 | 488 | 12 | 493 |
| BAD99527.1 | 1.95e-218 | 9 | 488 | 10 | 490 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1WKY_A | 5.63e-220 | 25 | 488 | 2 | 464 | Crystalstructure of alkaline mannanase from Bacillus sp. strain JAMB-602: catalytic domain and its Carbohydrate Binding Module [Bacillus sp. JAMB-602] |
| 3JUG_A | 5.34e-152 | 21 | 328 | 12 | 320 | Crystalstructure of endo-beta-1,4-mannanase from the alkaliphilic Bacillus sp. N16-5 [Bacillus sp. N16-5] |
| 2WHJ_A | 2.70e-150 | 31 | 333 | 1 | 306 | Understandinghow diverse mannanases recognise heterogeneous substrates [Salipaludibacillus agaradhaerens] |
| 2WHL_A | 3.32e-145 | 32 | 326 | 1 | 293 | Understandinghow diverse mannanases recognise heterogeneous substrates [Salipaludibacillus agaradhaerens] |
| 1BQC_A | 2.21e-79 | 30 | 327 | 1 | 302 | Beta-MannanaseFrom Thermomonospora Fusca [Thermobifida fusca],2MAN_A Mannotriose Complex Of Thermomonospora Fusca Beta-Mannanase [Thermobifida fusca],3MAN_A Mannohexaose Complex Of Thermomonospora Fusca Beta-mannanase [Thermobifida fusca] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| G1K3N4 | 1.48e-149 | 31 | 333 | 1 | 306 | Mannan endo-1,4-beta-mannosidase OS=Salipaludibacillus agaradhaerens OX=76935 PE=1 SV=1 |
| P51529 | 2.54e-85 | 15 | 334 | 21 | 343 | Mannan endo-1,4-beta-mannosidase OS=Streptomyces lividans OX=1916 GN=manA PE=1 SV=2 |
| B3PF24 | 3.62e-85 | 20 | 340 | 36 | 360 | Mannan endo-1,4-beta-mannosidase OS=Cellvibrio japonicus (strain Ueda107) OX=498211 GN=man5A PE=1 SV=1 |
| P22533 | 1.25e-62 | 52 | 319 | 55 | 326 | Beta-mannanase/endoglucanase A OS=Caldicellulosiruptor saccharolyticus OX=44001 GN=manA PE=1 SV=2 |
| P16699 | 2.54e-33 | 329 | 488 | 353 | 513 | Mannan endo-1,4-beta-mannosidase A and B OS=Caldalkalibacillus mannanilyticus (strain DSM 16130 / CIP 109019 / JCM 10596 / AM-001) OX=1236954 PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
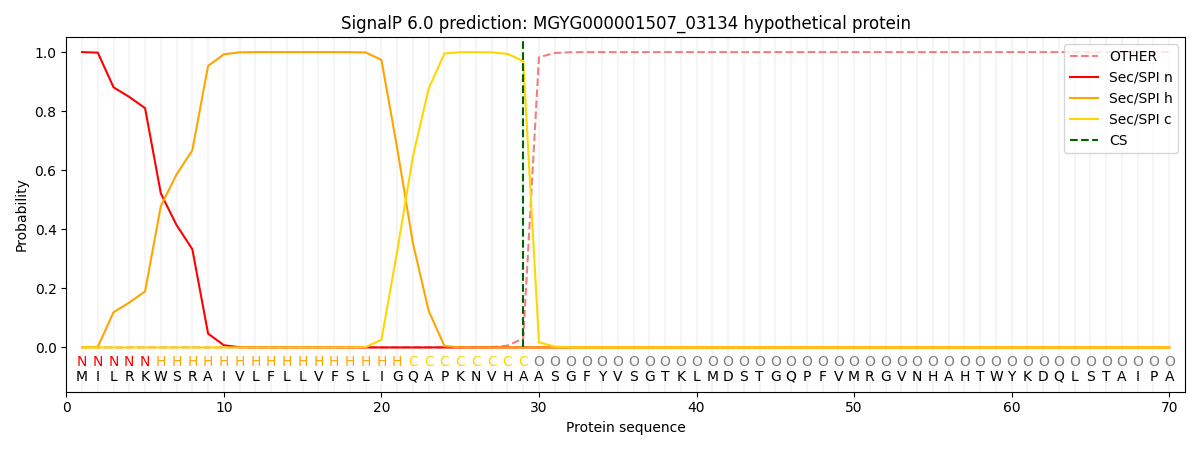
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000235 | 0.999035 | 0.000221 | 0.000162 | 0.000161 | 0.000141 |
