You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001507_05444
You are here: Home > Sequence: MGYG000001507_05444
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Paenibacillus ihuae | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes; Bacilli; Paenibacillales; Paenibacillaceae; Paenibacillus; Paenibacillus ihuae | |||||||||||
| CAZyme ID | MGYG000001507_05444 | |||||||||||
| CAZy Family | GT2 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 82714; End: 83622 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GT2 | 5 | 131 | 1.2e-16 | 0.7235294117647059 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd00761 | Glyco_tranf_GTA_type | 3.50e-18 | 6 | 128 | 1 | 118 | Glycosyltransferase family A (GT-A) includes diverse families of glycosyl transferases with a common GT-A type structural fold. Glycosyltransferases (GTs) are enzymes that synthesize oligosaccharides, polysaccharides, and glycoconjugates by transferring the sugar moiety from an activated nucleotide-sugar donor to an acceptor molecule, which may be a growing oligosaccharide, a lipid, or a protein. Based on the stereochemistry of the donor and acceptor molecules, GTs are classified as either retaining or inverting enzymes. To date, all GT structures adopt one of two possible folds, termed GT-A fold and GT-B fold. This hierarchy includes diverse families of glycosyl transferases with a common GT-A type structural fold, which has two tightly associated beta/alpha/beta domains that tend to form a continuous central sheet of at least eight beta-strands. The majority of the proteins in this superfamily are Glycosyltransferase family 2 (GT-2) proteins. But it also includes families GT-43, GT-6, GT-8, GT13 and GT-7; which are evolutionarily related to GT-2 and share structure similarities. |
| pfam00535 | Glycos_transf_2 | 1.44e-12 | 5 | 124 | 1 | 114 | Glycosyl transferase family 2. Diverse family, transferring sugar from UDP-glucose, UDP-N-acetyl- galactosamine, GDP-mannose or CDP-abequose, to a range of substrates including cellulose, dolichol phosphate and teichoic acids. |
| pfam13641 | Glyco_tranf_2_3 | 2.61e-09 | 4 | 215 | 4 | 198 | Glycosyltransferase like family 2. Members of this family of prokaryotic proteins include putative glucosyltransferase, which are involved in bacterial capsule biosynthesis. |
| COG0463 | WcaA | 4.02e-08 | 1 | 105 | 2 | 101 | Glycosyltransferase involved in cell wall bisynthesis [Cell wall/membrane/envelope biogenesis]. |
| cd06420 | GT2_Chondriotin_Pol_N | 5.34e-08 | 6 | 188 | 1 | 140 | N-terminal domain of Chondroitin polymerase functions as a GalNAc transferase. Chondroitin polymerase is a two domain, bi-functional protein. The N-terminal domain functions as a GalNAc transferase. The bacterial chondroitin polymerase catalyzes elongation of the chondroitin chain by alternatively transferring the GlcUA and GalNAc moiety from UDP-GlcUA and UDP-GalNAc to the non-reducing ends of the chondroitin chain. The enzyme consists of N-terminal and C-terminal domains in which the two active sites catalyze the addition of GalNAc and GlcUA, respectively. Chondroitin chains range from 40 to over 100 repeating units of the disaccharide. Sulfated chondroitins are involved in the regulation of various biological functions such as central nervous system development, wound repair, infection, growth factor signaling, and morphogenesis, in addition to its conventional structural roles. In Caenorhabditis elegans, chondroitin is an essential factor for the worm to undergo cytokinesis and cell division. Chondroitin is synthesized as proteoglycans, sulfated and secreted to the cell surface or extracellular matrix. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QSF45363.1 | 1.43e-219 | 1 | 302 | 1 | 302 |
| AIQ15488.1 | 1.56e-200 | 1 | 301 | 1 | 300 |
| AIQ66475.1 | 5.20e-199 | 1 | 301 | 1 | 300 |
| QQZ62854.1 | 2.12e-198 | 1 | 301 | 1 | 300 |
| CQR51605.1 | 2.12e-198 | 1 | 301 | 1 | 300 |
Swiss-Prot Hits help
SignalP and Lipop Annotations help
This protein is predicted as OTHER
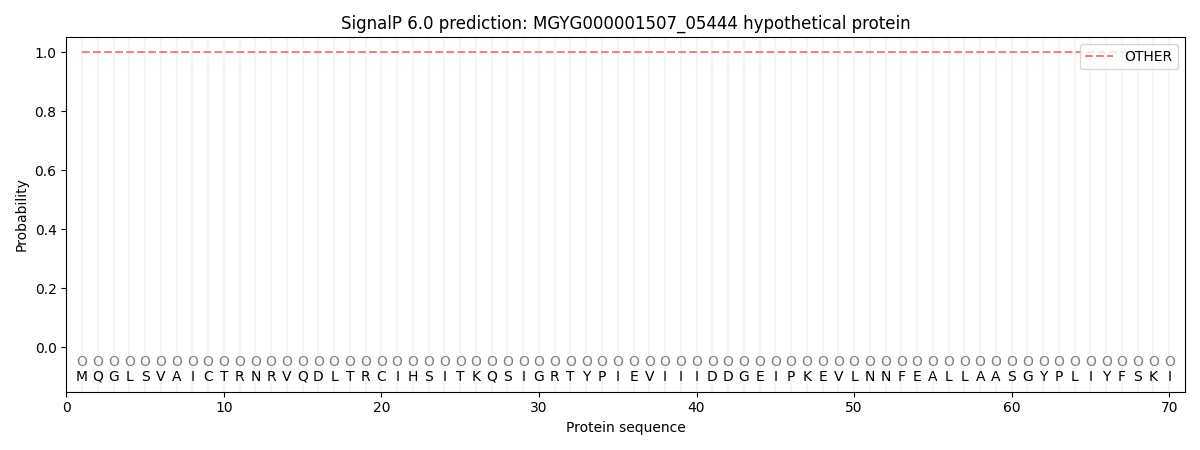
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000068 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
