You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001514_04807
You are here: Home > Sequence: MGYG000001514_04807
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Paenibacillus_A ihumii | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes; Bacilli; Paenibacillales; Paenibacillaceae; Paenibacillus_A; Paenibacillus_A ihumii | |||||||||||
| CAZyme ID | MGYG000001514_04807 | |||||||||||
| CAZy Family | CE12 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 3600380; End: 3601198 Strand: + | |||||||||||
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG0657 | Aes | 1.00e-21 | 52 | 264 | 79 | 310 | Acetyl esterase/lipase [Lipid transport and metabolism]. |
| COG1506 | DAP2 | 1.47e-16 | 101 | 264 | 454 | 616 | Dipeptidyl aminopeptidase/acylaminoacyl peptidase [Amino acid transport and metabolism]. |
| pfam00326 | Peptidase_S9 | 1.05e-11 | 96 | 264 | 31 | 209 | Prolyl oligopeptidase family. |
| pfam07859 | Abhydrolase_3 | 1.25e-09 | 101 | 240 | 49 | 206 | alpha/beta hydrolase fold. This catalytic domain is found in a very wide range of enzymes. |
| COG0412 | DLH | 3.70e-04 | 47 | 265 | 22 | 234 | Dienelactone hydrolase [Secondary metabolites biosynthesis, transport and catabolism]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| ASR46787.1 | 9.70e-118 | 1 | 268 | 1 | 250 |
| AUS27537.1 | 2.75e-117 | 1 | 268 | 1 | 250 |
| QPK59380.1 | 2.75e-117 | 1 | 267 | 1 | 249 |
| AIY07762.1 | 2.75e-117 | 1 | 268 | 1 | 250 |
| AHM66957.1 | 2.75e-117 | 1 | 268 | 1 | 250 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6A6O_A | 1.17e-72 | 47 | 262 | 49 | 277 | ChainA, Esterase/lipase-like protein [Caldicellulosiruptor lactoaceticus 6A] |
| 3HXK_A | 1.12e-40 | 53 | 263 | 44 | 264 | CrystalStructure of a sugar hydrolase (YeeB) from Lactococcus lactis, Northeast Structural Genomics Consortium Target KR108 [Lactococcus lactis subsp. lactis],3HXK_B Crystal Structure of a sugar hydrolase (YeeB) from Lactococcus lactis, Northeast Structural Genomics Consortium Target KR108 [Lactococcus lactis subsp. lactis],3HXK_C Crystal Structure of a sugar hydrolase (YeeB) from Lactococcus lactis, Northeast Structural Genomics Consortium Target KR108 [Lactococcus lactis subsp. lactis],3HXK_D Crystal Structure of a sugar hydrolase (YeeB) from Lactococcus lactis, Northeast Structural Genomics Consortium Target KR108 [Lactococcus lactis subsp. lactis] |
| 4Q3K_A | 1.13e-35 | 39 | 263 | 41 | 256 | Crystalstructure of MGS-M1, an alpha/beta hydrolase enzyme from a Medee basin deep-sea metagenome library [unidentified],4Q3K_B Crystal structure of MGS-M1, an alpha/beta hydrolase enzyme from a Medee basin deep-sea metagenome library [unidentified] |
| 3BJR_A | 1.39e-35 | 53 | 261 | 51 | 279 | ChainA, Putative carboxylesterase [Lactiplantibacillus plantarum WCFS1] |
| 7DWC_A | 1.22e-30 | 55 | 261 | 55 | 261 | ChainA, Xylanase [Bacteroides thetaiotaomicron VPI-5482],7DWC_B Chain B, Xylanase [Bacteroides thetaiotaomicron VPI-5482],7DWC_C Chain C, Xylanase [Bacteroides thetaiotaomicron VPI-5482],7DWC_D Chain D, Xylanase [Bacteroides thetaiotaomicron VPI-5482] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P26223 | 2.40e-34 | 52 | 246 | 402 | 612 | Endo-1,4-beta-xylanase B OS=Butyrivibrio fibrisolvens OX=831 GN=xynB PE=3 SV=1 |
| D5EV35 | 7.44e-30 | 55 | 261 | 50 | 263 | Acetylxylan esterase OS=Prevotella ruminicola (strain ATCC 19189 / JCM 8958 / 23) OX=264731 GN=axeA1 PE=1 SV=1 |
| Q50681 | 3.61e-07 | 38 | 264 | 167 | 419 | Probable carboxylic ester hydrolase LipM OS=Mycobacterium tuberculosis (strain ATCC 25618 / H37Rv) OX=83332 GN=lipM PE=1 SV=1 |
| A0A097ZPE0 | 7.20e-06 | 37 | 164 | 2112 | 2253 | Non-reducing polyketide synthase andM OS=Emericella variicolor OX=1549217 GN=andM PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
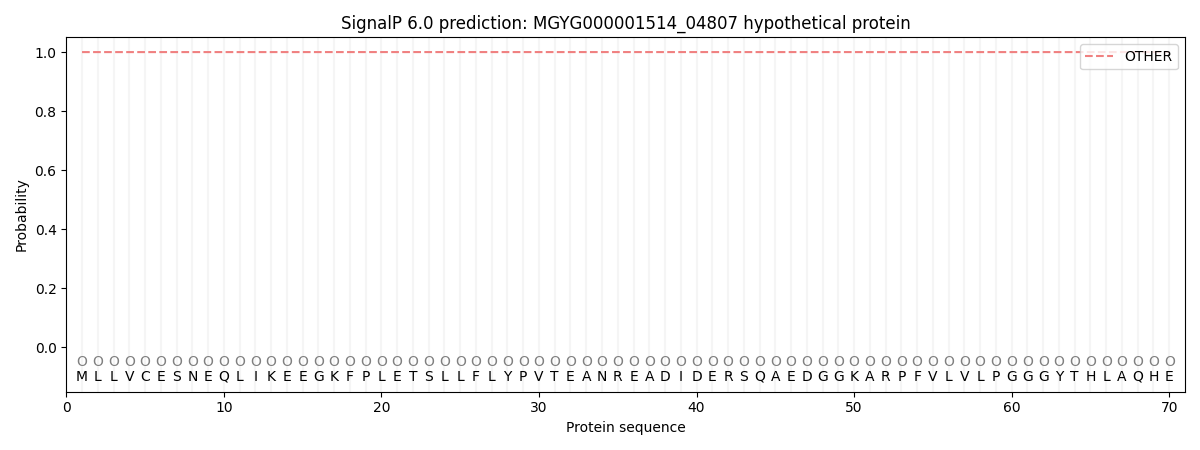
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000055 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
