You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000001526_03987
You are here: Home > Sequence: MGYG000001526_03987
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
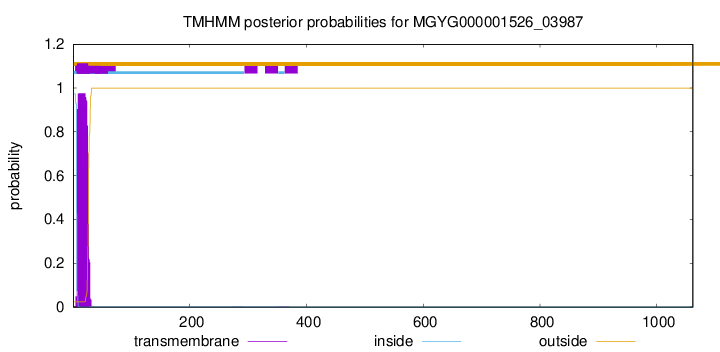
TMHMM annotations
Basic Information help
| Species | Paenibacillus_A senegalimassiliensis | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes; Bacilli; Paenibacillales; Paenibacillaceae; Paenibacillus_A; Paenibacillus_A senegalimassiliensis | |||||||||||
| CAZyme ID | MGYG000001526_03987 | |||||||||||
| CAZy Family | GH28 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 1196995; End: 1200183 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH28 | 251 | 589 | 2.4e-39 | 0.9230769230769231 |
| CBM66 | 902 | 1056 | 6.2e-30 | 0.9870967741935484 |
| CBM66 | 624 | 775 | 1e-26 | 0.967741935483871 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG5434 | Pgu1 | 1.99e-21 | 221 | 603 | 82 | 453 | Polygalacturonase [Carbohydrate transport and metabolism]. |
| pfam06439 | DUF1080 | 9.12e-12 | 618 | 775 | 6 | 178 | Domain of Unknown Function (DUF1080). This family has structural similarity to an endo-1,3-1,4-beta glucanase belonging to glycoside hydrolase family 16. However, the structure surrounding the active site differs from that of the endo-1,3-1,4-beta glucanase. |
| cd11304 | Cadherin_repeat | 1.88e-11 | 794 | 883 | 5 | 97 | Cadherin tandem repeat domain. Cadherins are glycoproteins involved in Ca2+-mediated cell-cell adhesion. The cadherin repeat domains occur as tandem repeats in the extracellular regions, which are thought to mediate cell-cell contact when bound to calcium. They play numerous roles in cell fate, signalling, proliferation, differentiation, and migration; members include E-, N-, P-, T-, VE-, CNR-, proto-, and FAT-family cadherin, desmocollin, and desmoglein, a large variety of domain architectures with varying repeat copy numbers. Cadherin-repeat containing proteins exist as monomers, homodimers, or heterodimers. |
| pfam06439 | DUF1080 | 3.08e-10 | 891 | 1059 | 3 | 182 | Domain of Unknown Function (DUF1080). This family has structural similarity to an endo-1,3-1,4-beta glucanase belonging to glycoside hydrolase family 16. However, the structure surrounding the active site differs from that of the endo-1,3-1,4-beta glucanase. |
| smart00112 | CA | 3.73e-06 | 809 | 885 | 1 | 80 | Cadherin repeats. Cadherins are glycoproteins involved in Ca2+-mediated cell-cell adhesion. Cadherin domains occur as repeats in the extracellular regions which are thought to mediate cell-cell contact when bound to calcium. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| ACX65654.1 | 0.0 | 83 | 1058 | 36 | 1018 |
| QOT12100.1 | 0.0 | 3 | 1058 | 5 | 1018 |
| QUL54880.1 | 0.0 | 111 | 1060 | 71 | 1050 |
| CQR58237.1 | 7.17e-147 | 96 | 596 | 20 | 510 |
| AWV34615.1 | 2.79e-146 | 104 | 596 | 29 | 510 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2UVE_A | 1.69e-09 | 216 | 333 | 151 | 273 | Structureof Yersinia enterocolitica Family 28 Exopolygalacturonase [Yersinia enterocolitica],2UVE_B Structure of Yersinia enterocolitica Family 28 Exopolygalacturonase [Yersinia enterocolitica],2UVF_A Structure of Yersinia enterocolitica Family 28 Exopolygalacturonase in Complex with Digalaturonic Acid [Yersinia enterocolitica],2UVF_B Structure of Yersinia enterocolitica Family 28 Exopolygalacturonase in Complex with Digalaturonic Acid [Yersinia enterocolitica] |
| 5OLP_A | 3.86e-09 | 229 | 430 | 50 | 276 | Galacturonidase[Bacteroides thetaiotaomicron VPI-5482],5OLP_B Galacturonidase [Bacteroides thetaiotaomicron VPI-5482] |
| 4MXN_A | 2.73e-07 | 235 | 337 | 33 | 127 | Crystalstructure of a putative glycosyl hydrolase (PARMER_00599) from Parabacteroides merdae ATCC 43184 at 1.95 A resolution [Parabacteroides merdae ATCC 43184],4MXN_B Crystal structure of a putative glycosyl hydrolase (PARMER_00599) from Parabacteroides merdae ATCC 43184 at 1.95 A resolution [Parabacteroides merdae ATCC 43184],4MXN_C Crystal structure of a putative glycosyl hydrolase (PARMER_00599) from Parabacteroides merdae ATCC 43184 at 1.95 A resolution [Parabacteroides merdae ATCC 43184],4MXN_D Crystal structure of a putative glycosyl hydrolase (PARMER_00599) from Parabacteroides merdae ATCC 43184 at 1.95 A resolution [Parabacteroides merdae ATCC 43184] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| P15922 | 3.06e-09 | 219 | 610 | 149 | 579 | Exo-poly-alpha-D-galacturonosidase OS=Dickeya chrysanthemi OX=556 GN=pehX PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
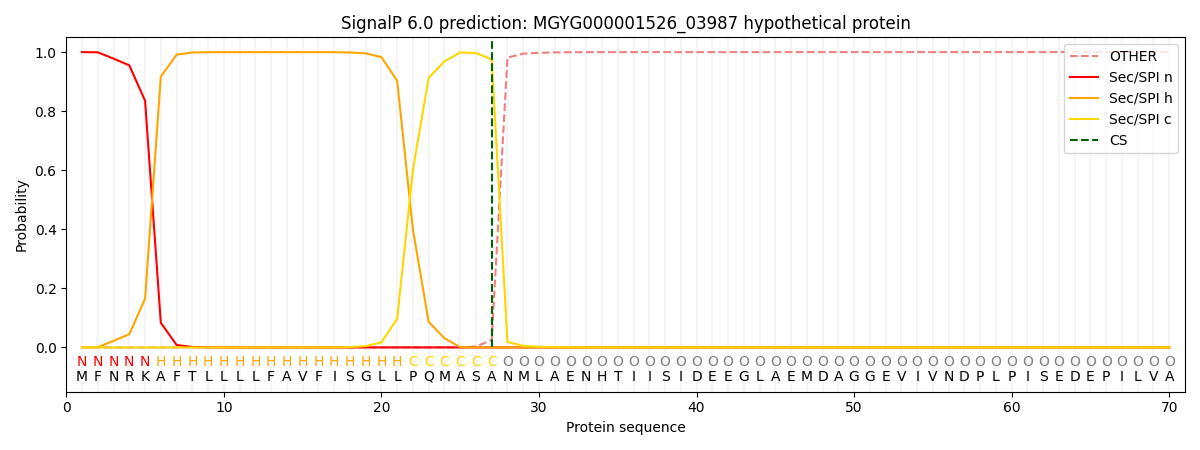
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000347 | 0.998815 | 0.000246 | 0.000202 | 0.000191 | 0.000164 |